Identification and genetic characterization of PmrA-regulated genes and genes involved in polymyxin B resistance in Salmonella enterica serovar typhimurium
- PMID: 12438352
- PMCID: PMC133008
- DOI: 10.1128/IAI.70.12.6770-6778.2002
Identification and genetic characterization of PmrA-regulated genes and genes involved in polymyxin B resistance in Salmonella enterica serovar typhimurium
Abstract
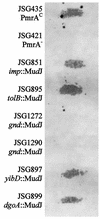
Salmonella enterica serovar Typhimurium encounters antimicrobial peptides (AP) within the phagosomes of professional phagocytes and at intestinal mucosal surfaces. Salmonella serovar Typhimurium utilizes the two-component regulatory system PmrA-PmrB, which is activated in response to the environmental conditions encountered in vivo, to regulate resistance to several AP, including polymyxin B (PM). Random MudJ transposon mutagenesis was used to identify PmrA-PmrB-regulated genes, as well as genetic loci necessary for PM resistance. Three different phenotypic classes of genes were identified: those necessary for PM resistance and regulated by PmrA, those necessary for PM resistance and not regulated by PmrA, and PmrA-regulated genes not required for PM resistance. Loci identified as necessary for PM resistance showed between 6- and 192-fold increased sensitivities to PM, and transposon insertion sites include surA, tolB, and gnd. PmrA-regulated loci identified included dgoA and yibD and demonstrated 500- and 2,500-fold activation by PmrA, respectively. The role of the identified loci in aminoarabinose modification of lipid A was determined by paper chromatography. The gnd mutant demonstrated a loss of aminoarabinose from lipid A, which was suggested to be due to a polar effect on the downstream gene pmrE. The remaining PM(s) mutants (surA and tolB), as well as the two PmrA-regulated gene (yibD and dgoA) mutants, retained aminoarabinose on lipid A. yibD, dgoA, and gnd (likely affecting pmrE) played no role in PmrA-regulated resistance to high iron concentrations, while surA and tolB mutations grew poorly on high iron media. All PM(s) mutants identified in this study demonstrated a defect in virulence compared to wild-type Salmonella serovar Typhimurium when administered orally to mice, while the PmrA-regulated gene (yibD and dgoA) mutants showed normal virulence in mice. These data broaden our understanding of in vivo gene regulation, lipopolysaccharide modification, and mechanisms of resistance to AP in enteric bacteria.
Figures


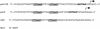
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

