Expression of a novel Leishmania gene encoding a histone H1-like protein in Leishmania major modulates parasite infectivity in vitro
- PMID: 12438377
- PMCID: PMC132950
- DOI: 10.1128/IAI.70.12.6976-6986.2002
Expression of a novel Leishmania gene encoding a histone H1-like protein in Leishmania major modulates parasite infectivity in vitro
Abstract
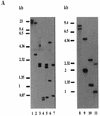
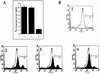
We describe identification and characterization of a novel two-copy gene of the parasitic protozoan Leishmania that encodes a nuclear protein designated LNP18. This protein is highly conserved in the genus Leishmania, and it is developmentally regulated. It is an alanine- and lysine-rich protein with potential bipartite nuclear targeting sequence sites. LNP18 shows sequence similarity to H1 histones of trypanosomatids and of higher eukaryotes and in particular with histone H1 of Leishmania major. The nuclear localization of LNP18 was determined by indirect immunofluorescence and Western blot analysis of isolated nuclei by using antibodies raised against the recombinant protein as probes. The antibodies recognized predominantly a 18-kDa band or a 18-kDa-16-kDa doublet. Photochemical cross-linking of intact parasites followed by Western blot analysis provided evidence that LNP18 is indeed a DNA-binding protein. Generation of transfectants overexpressing LNP18 allowed us to determine the role of this protein in Leishmania infection of macrophages in vitro. These studies revealed that transfectants overexpressing LNP18 are significantly less infective than transfectants with the vector alone and suggested that the level of LNP18 expression modulates Leishmania infectivity, as assessed in vitro.
Figures



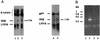
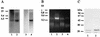

Similar articles
-
Histone H1 expression varies during the Leishmania major life cycle.Mol Biochem Parasitol. 1997 Feb;84(2):215-27. doi: 10.1016/s0166-6851(96)02801-0. Mol Biochem Parasitol. 1997. PMID: 9084041
-
Leishmania histone H1 overexpression delays parasite cell-cycle progression, parasite differentiation and reduces Leishmania infectivity in vivo.Mol Microbiol. 2006 Jun;60(6):1457-73. doi: 10.1111/j.1365-2958.2006.05205.x. Mol Microbiol. 2006. PMID: 16796681
-
Cloning and characterization of a Golgi-associated GTP-binding protein homologue from Leishmania major.Mol Biochem Parasitol. 1993 Nov;62(1):73-82. doi: 10.1016/0166-6851(93)90179-2. Mol Biochem Parasitol. 1993. PMID: 8114828
-
Leishmania major: histone H1 gene expression from the sw3 locus.Exp Parasitol. 1999 Feb;91(2):151-60. doi: 10.1006/expr.1998.4378. Exp Parasitol. 1999. PMID: 9990343
-
Canonical and variant histones of protozoan parasites.Front Biosci (Landmark Ed). 2011 Jun 1;16(6):2086-105. doi: 10.2741/3841. Front Biosci (Landmark Ed). 2011. PMID: 21622164 Review.
Cited by
-
Genomic Analysis of Colombian Leishmania panamensis strains with different level of virulence.Sci Rep. 2018 Nov 26;8(1):17336. doi: 10.1038/s41598-018-35778-6. Sci Rep. 2018. PMID: 30478412 Free PMC article.
-
Molecular characterization and immune protection of an AN1-like zinc finger protein of Eimeria tenella.Parasitol Res. 2020 Feb;119(2):623-635. doi: 10.1007/s00436-019-06545-x. Epub 2019 Nov 22. Parasitol Res. 2020. PMID: 31758298
-
The high mobility group protein HMO1 functions as a linker histone in yeast.Epigenetics Chromatin. 2016 Mar 30;9:13. doi: 10.1186/s13072-016-0062-8. eCollection 2016. Epigenetics Chromatin. 2016. PMID: 27030801 Free PMC article.
-
An inhibitor-driven study for enhancing the selectivity of indirubin derivatives towards leishmanial Glycogen Synthase Kinase-3 over leishmanial cdc2-related protein kinase 3.Parasit Vectors. 2014 May 20;7:234. doi: 10.1186/1756-3305-7-234. Parasit Vectors. 2014. PMID: 24886176 Free PMC article.
-
H2B.V demarcates divergent strand-switch regions, some tDNA loci, and genome compartments in Trypanosoma cruzi and affects parasite differentiation and host cell invasion.PLoS Pathog. 2022 Feb 18;18(2):e1009694. doi: 10.1371/journal.ppat.1009694. eCollection 2022 Feb. PLoS Pathog. 2022. PMID: 35180281 Free PMC article.
References
-
- Aslund, L., L. Carisson, J. Henriksson, M. Rydaker, C. Toro, N. Galanti, and U. Pettersson. 1994. A gene family encoding heterogeneous histone H1 proteins in Trypanosoma cruzi. Mol. Biochem. Parasitol. 65:317-330. - PubMed
-
- Astolfi, F. S., C. Martins de Sa, and E. S. Gander. 1980. On the chromatin structure of Trypanosoma cruzi. Mol. Biochem. Parasitol. 1:45-53. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

