RIP4 (DIK/PKK), a novel member of the RIP kinase family, activates NF-kappa B and is processed during apoptosis
- PMID: 12446564
- PMCID: PMC1308326
- DOI: 10.1093/embo-reports/kvf236
RIP4 (DIK/PKK), a novel member of the RIP kinase family, activates NF-kappa B and is processed during apoptosis
Abstract
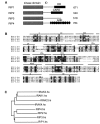
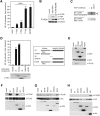
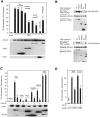
RIP1 and its homologs, RIP2 and RIP3, form part of a family of Ser/Thr kinases that regulate signal transduction processes leading to NF-kappa B activation. Here, we identify RIP4 (DIK/PKK) as a novel member of the RIP kinase family. RIP4 contains an N-terminal RIP-like kinase domain and a C-terminal region characterized by the presence of 11 ankyrin repeats. Overexpression of RIP4 leads to activation of NF-kappa B and JNK. Kinase inactive RIP4 or a truncated version containing the ankyrin repeats have a dominant negative (DN) effect on NF-kappa B induction by multiple stimuli. RIP4 binds to several members of the TRAF protein family, and DN versions of TRAF1, TRAF3 and TRAF6 inhibit RIP4-induced NF-kappa B activation. Moreover, RIP4 is cleaved after Asp340 and Asp378 during Fas-induced apoptosis. These data suggest that RIP4 is involved in NF-kappa B and JNK signaling and that caspase-dependent processing of RIP4 may negatively regulate NF-kappa B-dependent pro-survival or pro-inflammatory signals.
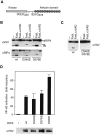
Figures
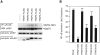
Similar articles
-
Protein kinase C-associated kinase (PKK) mediates Bcl10-independent NF-kappa B activation induced by phorbol ester.J Biol Chem. 2002 Aug 30;277(35):31871-6. doi: 10.1074/jbc.M202222200. Epub 2002 Jun 28. J Biol Chem. 2002. PMID: 12091384
-
RIP2 is a novel NF-kappaB-activating and cell death-inducing kinase.J Biol Chem. 1998 Jul 3;273(27):16968-75. doi: 10.1074/jbc.273.27.16968. J Biol Chem. 1998. PMID: 9642260
-
A catalytically inactive form of protein kinase C-associated kinase/receptor interacting protein 4, a protein kinase C beta-associated kinase that mediates NF-kappa B activation, interferes with early B cell development.J Immunol. 2003 Aug 15;171(4):1875-80. doi: 10.4049/jimmunol.171.4.1875. J Immunol. 2003. PMID: 12902489
-
Identification of CARDIAK, a RIP-like kinase that associates with caspase-1.Curr Biol. 1998 Jul 16;8(15):885-8. doi: 10.1016/s0960-9822(07)00352-1. Curr Biol. 1998. PMID: 9705938
-
The RIP kinases: crucial integrators of cellular stress.Trends Biochem Sci. 2005 Mar;30(3):151-9. doi: 10.1016/j.tibs.2005.01.003. Trends Biochem Sci. 2005. PMID: 15752987 Review.
Cited by
-
Role of Receptor Interacting Protein (RIP) kinases in cancer.Genes Dis. 2021 Nov 18;9(6):1579-1593. doi: 10.1016/j.gendis.2021.10.007. eCollection 2022 Nov. Genes Dis. 2021. PMID: 36157481 Free PMC article. Review.
-
Overexpression of RIPK4 Predicts Poor Prognosis and Promotes Metastasis in Ovarian Cancer.Biomed Res Int. 2021 Jun 2;2021:6622439. doi: 10.1155/2021/6622439. eCollection 2021. Biomed Res Int. 2021. PMID: 34124253 Free PMC article.
-
RIPK protein kinase family: Atypical lives of typical kinases.Semin Cell Dev Biol. 2021 Jan;109:96-105. doi: 10.1016/j.semcdb.2020.06.014. Epub 2020 Jul 27. Semin Cell Dev Biol. 2021. PMID: 32732131 Free PMC article. Review.
-
Pathogen-mediated proteolysis of the cell death regulator RIPK1 and the host defense modulator RIPK2 in human aortic endothelial cells.PLoS Pathog. 2012;8(6):e1002723. doi: 10.1371/journal.ppat.1002723. Epub 2012 Jun 7. PLoS Pathog. 2012. PMID: 22685397 Free PMC article.
-
The ANKK1 protein associated with addictions has nuclear and cytoplasmic localization and shows a differential response of Ala239Thr to apomorphine.Neurotox Res. 2011 Jul;20(1):32-9. doi: 10.1007/s12640-010-9219-6. Epub 2010 Sep 16. Neurotox Res. 2011. PMID: 20845092
References
-
- Bähr C., Rohwer A., Stempka L., Rincke G., Marks F. and Gschwendt M. (2000) DIK, a novel protein kinase that interacts with protein kinase Cδ. Cloning, characterization, and gene analysis. J. Biol. Chem., 275, 36350–36357. - PubMed
-
- Bucher P., Karplus K., Moeri N. and Hofmann K. (1996) A flexible search technique based on generalized profiles. Comp. Chem., 20, 3–24. - PubMed
-
- Chen L., Haider K., Ponda M., Cariappa A., Rowitch D. and Pillai S. (2001) Protein kinase C-associated kinase (PKK), a novel membrane-associated, ankyrin repeat-containing protein kinase. J. Biol. Chem., 276, 21737–21744. - PubMed
-
- Chin A.I., Dempsey P.W., Bruhn K., Miller J.F., Xu Y. and Cheng G. (2002) Involvement of receptor-interacting protein 2 in innate and adaptive immune responses. Nature, 416, 190–194. - PubMed
-
- Hanks S.K. and Hunter T. (1995) Protein kinases 6. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification. FASEB J., 9, 576–596. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

