Role of ubiquitination in retro-translocation of cholera toxin and escape of cytosolic degradation
- PMID: 12446567
- PMCID: PMC1308323
- DOI: 10.1093/embo-reports/kvf239
Role of ubiquitination in retro-translocation of cholera toxin and escape of cytosolic degradation
Abstract
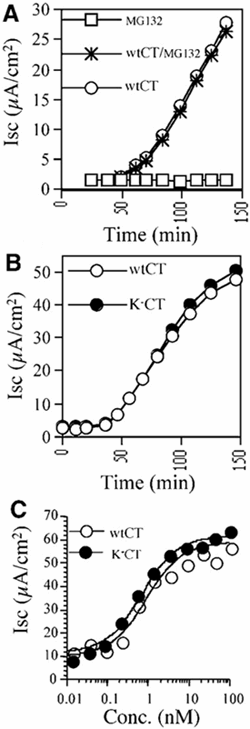
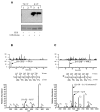
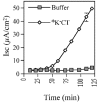
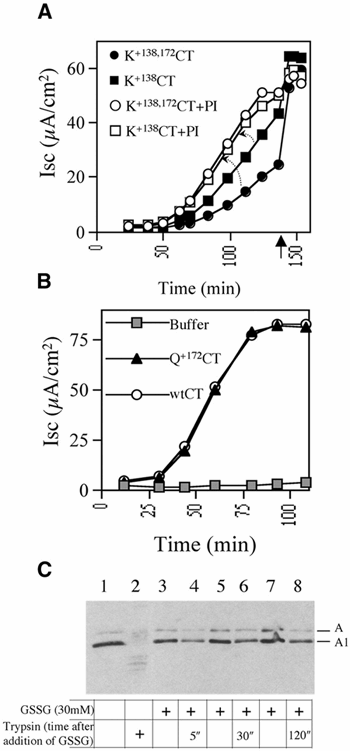
Cholera toxin travels from the cell surface of affected mammalian cells to the endoplasmic reticulum (ER), where the A1 chain is released and retro-translocated across the ER membrane into the cytosol. We have tested whether, as in other cases, retro-translocation requires poly-ubiquitination. We show that an A1 chain mutant that lacks lysines and has a blocked N-terminus, and therefore cannot be ubiquitinated, remains active in vivo. The A1 chain is not degraded in the cytosol, as demonstrated by the fact that proteasome inhibitors do not stimulate its activity. When additional lysines are introduced into the A1 chain, moderate degradation by the proteasome is observed. The unfolded A1 chain rapidly refolds in vitro. These results show that poly-ubiquitination is not required for retro-translocation of all proteins across the ER membrane and indicate that the reason why the toxin escapes degradation in the cytosol may be both its paucity of lysines and its rapid refolding.
Figures
References
-
- Deeks E.D., Cook J.P., Day P.J., Smith D.C., Roberts L.M. and Lord J.M. (2002) The low lysine content of ricin A chain reduces the risk of proteolytic degradation after translocation from the endoplasmic reticulum to the cytosol. Biochemistry, 41, 3405–3413. - PubMed
-
- Hazes B. and Read R.J. (1997) Accumulating evidence suggests that several AB-toxins subvert the endoplasmic reticulum-associated protein degradation pathway to enter target cells. Biochemistry, 36, 11051–11054. - PubMed
-
- Hershko A. and Ciechanover A. (1998) The ubiquitin system. Annu. Rev. Biochem., 67, 425–479. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

