Role of transcription in plasmid maintenance in the hpr1Delta mutant of Saccharomyces cerevisiae
- PMID: 12446793
- PMCID: PMC139893
- DOI: 10.1128/MCB.22.24.8763-8773.2002
Role of transcription in plasmid maintenance in the hpr1Delta mutant of Saccharomyces cerevisiae
Abstract
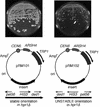
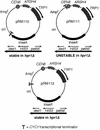
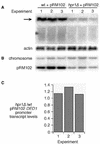
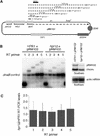
The Saccharomyces cerevisiae hyperrecombination mutation hpr1Delta results in instability of sequences between direct repeats that is dependent on transcription of the repeat. Here it is shown that the HPR1 gene also functions in plasmid stability in the presence of destabilizing transcription elongation. In the hpr1Delta mutant, plasmid instability results from unchecked transcription elongation, which can be suppressed by a strong transcription terminator. The plasmid system has been used to examine in vivo aspects of transcription in the absence of Hpr1p. Nuclear run-on studies suggest that there is an increased RNA polymerase II density in the hpr1Delta mutant strain, but this is not accompanied by an increase in accumulation of cytoplasmic mRNA. Suppression of plasmid instability in hpr1Delta can also be achieved by high-copy expression of the RNA splicing factor SUB2, which has recently been proposed to function in mRNA export, in addition to its role in pre-mRNA splicing. High-copy-number SUB2 expression is accompanied by an increase in message accumulation from the plasmid, suggesting that the mechanism of suppression by Sub2p involves the formation of mature mRNA. Models for the role of Hpr1p in mature mRNA formation and the cause of plasmid instability in the absence of the Hpr1 protein are discussed.
Figures


Similar articles
-
High-copy-number expression of Sub2p, a member of the RNA helicase superfamily, suppresses hpr1-mediated genomic instability.Mol Cell Biol. 2001 Aug;21(16):5459-70. doi: 10.1128/MCB.21.16.5459-5470.2001. Mol Cell Biol. 2001. PMID: 11463828 Free PMC article.
-
hpr1Delta affects ribosomal DNA recombination and cell life span in Saccharomyces cerevisiae.Mol Cell Biol. 2002 Jan;22(2):421-9. doi: 10.1128/MCB.22.2.421-429.2002. Mol Cell Biol. 2002. PMID: 11756539 Free PMC article.
-
Stable mRNP formation and export require cotranscriptional recruitment of the mRNA export factors Yra1p and Sub2p by Hpr1p.Mol Cell Biol. 2002 Dec;22(23):8241-53. doi: 10.1128/MCB.22.23.8241-8253.2002. Mol Cell Biol. 2002. PMID: 12417727 Free PMC article.
-
Tho1, a novel hnRNP, and Sub2 provide alternative pathways for mRNP biogenesis in yeast THO mutants.Mol Cell Biol. 2006 Jun;26(12):4387-98. doi: 10.1128/MCB.00234-06. Mol Cell Biol. 2006. PMID: 16738307 Free PMC article.
-
The 2 micron plasmid of Saccharomyces cerevisiae: a miniaturized selfish genome with optimized functional competence.Plasmid. 2013 Jul;70(1):2-17. doi: 10.1016/j.plasmid.2013.03.001. Epub 2013 Mar 27. Plasmid. 2013. PMID: 23541845 Review.
Cited by
-
HYPER RECOMBINATION1 of the THO/TREX complex plays a role in controlling transcription of the REVERSION-TO-ETHYLENE SENSITIVITY1 gene in Arabidopsis.PLoS Genet. 2015 Feb 13;11(2):e1004956. doi: 10.1371/journal.pgen.1004956. eCollection 2015 Feb. PLoS Genet. 2015. PMID: 25680185 Free PMC article.
-
Replication fork progression is impaired by transcription in hyperrecombinant yeast cells lacking a functional THO complex.Mol Cell Biol. 2006 Apr;26(8):3327-34. doi: 10.1128/MCB.26.8.3327-3334.2006. Mol Cell Biol. 2006. PMID: 16581804 Free PMC article.
References
-
- Brill, S. J., and R. Sternglanz. 1988. Transcription-dependent DNA supercoiling in yeast DNA topoisomerase mutants. Cell 54:403-411. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
