A whole-genome shotgun optical map of Yersinia pestis strain KIM
- PMID: 12450857
- PMCID: PMC134435
- DOI: 10.1128/AEM.68.12.6321-6331.2002
A whole-genome shotgun optical map of Yersinia pestis strain KIM
Abstract
Yersinia pestis is the causative agent of the bubonic, septicemic, and pneumonic plagues (also known as black death) and has been responsible for recurrent devastating pandemics throughout history. To further understand this virulent bacterium and to accelerate an ongoing sequencing project, two whole-genome restriction maps (XhoI and PvuII) of Y. pestis strain KIM were constructed using shotgun optical mapping. This approach constructs ordered restriction maps from randomly sheared individual DNA molecules directly extracted from cells. The two maps served different purposes; the XhoI map facilitated sequence assembly by providing a scaffold for high-resolution alignment, while the PvuII map verified genome sequence assembly. Our results show that such maps facilitated the closure of sequence gaps and, most importantly, provided a purely independent means for sequence validation. Given the recent advancements to the optical mapping system, increased resolution and throughput are enabling such maps to guide sequence assembly at a very early stage of a microbial sequencing project.
Figures
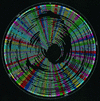
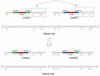




Similar articles
-
Whole-genome shotgun optical mapping of Rhodobacter sphaeroides strain 2.4.1 and its use for whole-genome shotgun sequence assembly.Genome Res. 2003 Sep;13(9):2142-51. doi: 10.1101/gr.1128803. Genome Res. 2003. PMID: 12952882 Free PMC article.
-
Whole-genome shotgun optical mapping of Rhodospirillum rubrum.Appl Environ Microbiol. 2005 Sep;71(9):5511-22. doi: 10.1128/AEM.71.9.5511-5522.2005. Appl Environ Microbiol. 2005. PMID: 16151144 Free PMC article.
-
Comparative scaffolding and gap filling of ancient bacterial genomes applied to two ancient Yersinia pestis genomes.Microb Genom. 2017 Jul 8;3(9):e000123. doi: 10.1099/mgen.0.000123. eCollection 2017 Sep. Microb Genom. 2017. PMID: 29114402 Free PMC article.
-
Yersinia pestis and the plague.Am J Clin Pathol. 2003 Jun;119 Suppl:S78-85. doi: 10.1309/DQM9-3R8Q-NQWB-FYU8. Am J Clin Pathol. 2003. PMID: 12951845 Review.
-
Genome and Evolution of Yersinia pestis.Adv Exp Med Biol. 2016;918:171-192. doi: 10.1007/978-94-024-0890-4_6. Adv Exp Med Biol. 2016. PMID: 27722863 Review.
Cited by
-
The genome of opportunistic fungal pathogen Fusarium oxysporum carries a unique set of lineage-specific chromosomes.Commun Biol. 2020 Jan 31;3(1):50. doi: 10.1038/s42003-020-0770-2. Commun Biol. 2020. PMID: 32005944 Free PMC article.
-
An algorithm for assembly of ordered restriction maps from single DNA molecules.Proc Natl Acad Sci U S A. 2006 Oct 24;103(43):15770-5. doi: 10.1073/pnas.0604040103. Epub 2006 Oct 16. Proc Natl Acad Sci U S A. 2006. PMID: 17043225 Free PMC article.
-
Rapid DNA mapping by fluorescent single molecule detection.Nucleic Acids Res. 2007;35(3):e16. doi: 10.1093/nar/gkl1044. Epub 2006 Dec 14. Nucleic Acids Res. 2007. PMID: 17175538 Free PMC article.
-
Error correcting optical mapping data.Gigascience. 2018 Jun 1;7(6):giy061. doi: 10.1093/gigascience/giy061. Gigascience. 2018. PMID: 29846578 Free PMC article.
-
Dynamics of genome rearrangement in bacterial populations.PLoS Genet. 2008 Jul 18;4(7):e1000128. doi: 10.1371/journal.pgen.1000128. PLoS Genet. 2008. PMID: 18650965 Free PMC article.
References
-
- Anantharaman, T. S., B. Mishra, and D. C. Schwartz. 1999. Genomics via optical mapping. III. Contiging genomic DNA and variations. The Seventh International Conference on Intelligent Systems for Molecular Biology, vol. 7, p. 18-27. - PubMed
-
- Anantharaman, T. S., B. Mishra, and D. C. Schwartz. 1998. Genomics via optical mapping. III. Contiging genomic DNA and variations. Courant Technical Report 760. Courant Institute, New York University, New York, NY.
-
- Anantharaman, T. S., B. Mishra, and D. C. Schwartz. 1997. Genomics via optical mapping. 2. Ordered restriction maps. J. Comput. Biol. 4:91-118. - PubMed
-
- Aston, C., C. Hiort, and D. C. Schwartz. 1999. Optical mapping: an approach for fine mapping. Methods Enzymol. 303:55-73. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

