Yeast Rpi1 is a putative transcriptional regulator that contributes to preparation for stationary phase
- PMID: 12455971
- PMCID: PMC118052
- DOI: 10.1128/EC.1.1.56-65.2002
Yeast Rpi1 is a putative transcriptional regulator that contributes to preparation for stationary phase
Abstract
The RPI1 gene of Saccharomyces cerevisiae was identified initially as a dosage suppressor of the heat shock sensitivity associated with overexpression of RAS2 (J. Kim and S. Powers, Mol. Cell. Biol. 11:3894-3904, 1991). Based on its failure to suppress mutationally activated RAS2, RPII was proposed to be a negative regulator of the Ras/cyclic AMP (cAMP) pathway that functions at a point upstream of Ras. We isolated RPI1 as a high-copy-number suppressor of the cell lysis defect associated with a null mutation in the MPK1 gene, which encodes the mitogen-activated protein kinase of the cell wall integrity-signaling pathway. Although the sequence of Rpil is not informative about its function, we present evidence that this protein resides in the nucleus, possesses a transcriptional activation domain, and affects the mRNA levels of several cell wall metabolism genes. In contrast to the previous report, we found that RPI1 overexpression suppresses defects associated with mutational hyperactivation of the Ras/cAMP pathway at all points including constitutive mutations in the cAMP-dependent protein kinase. We present additional genetic and biochemical evidence that Rpil functions independently of and in opposition to the Ras/cAMP pathway to promote preparations for the stationary phase. Among these preparations is a fortification of the cell wall that is antagonized by Ras pathway activity. This observation reveals a novel link between the Ras/cAMP pathway and cell wall integrity. Finally, we propose that inappropriate expression of RPI1 during log phase growth drives fortification of the cell wall and that this behavior is responsible for suppression of the mpkl cell lysis defect.
Figures

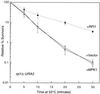





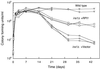


Similar articles
-
Overexpression of RPI1, a novel inhibitor of the yeast Ras-cyclic AMP pathway, down-regulates normal but not mutationally activated ras function.Mol Cell Biol. 1991 Aug;11(8):3894-904. doi: 10.1128/mcb.11.8.3894-3904.1991. Mol Cell Biol. 1991. PMID: 1649384 Free PMC article.
-
A family of genes required for maintenance of cell wall integrity and for the stress response in Saccharomyces cerevisiae.Proc Natl Acad Sci U S A. 1997 Dec 9;94(25):13804-9. doi: 10.1073/pnas.94.25.13804. Proc Natl Acad Sci U S A. 1997. PMID: 9391108 Free PMC article.
-
Saccharomyces cerevisiae Ras/cAMP pathway controls post-diauxic shift element-dependent transcription through the zinc finger protein Gis1.EMBO J. 2000 Jun 1;19(11):2569-79. doi: 10.1093/emboj/19.11.2569. EMBO J. 2000. PMID: 10835355 Free PMC article.
-
Novel sensing mechanisms and targets for the cAMP-protein kinase A pathway in the yeast Saccharomyces cerevisiae.Mol Microbiol. 1999 Sep;33(5):904-18. doi: 10.1046/j.1365-2958.1999.01538.x. Mol Microbiol. 1999. PMID: 10476026 Review.
-
Ras-regulated signaling processes in Saccharomyces cerevisiae.Curr Opin Genet Dev. 1991 Oct;1(3):370-7. doi: 10.1016/s0959-437x(05)80302-8. Curr Opin Genet Dev. 1991. PMID: 1668647 Review.
Cited by
-
A novel Ras inhibitor, Eri1, engages yeast Ras at the endoplasmic reticulum.Mol Cell Biol. 2003 Jul;23(14):4983-90. doi: 10.1128/MCB.23.14.4983-4990.2003. Mol Cell Biol. 2003. PMID: 12832483 Free PMC article.
-
Defects arising from whole-genome duplications in Saccharomyces cerevisiae.Genetics. 2004 Jul;167(3):1109-21. doi: 10.1534/genetics.104.029256. Genetics. 2004. PMID: 15280227 Free PMC article.
-
New Aspects of Invasive Growth Regulation Identified by Functional Profiling of MAPK Pathway Targets in Saccharomyces cerevisiae.Genetics. 2020 Sep;216(1):95-116. doi: 10.1534/genetics.120.303369. Epub 2020 Jul 14. Genetics. 2020. PMID: 32665277 Free PMC article.
-
Diverse conditions support near-zero growth in yeast: Implications for the study of cell lifespan.Microb Cell. 2019 Aug 20;6(9):397-413. doi: 10.15698/mic2019.09.690. Microb Cell. 2019. PMID: 31528631 Free PMC article. Review.
-
Gpi19, the Saccharomyces cerevisiae homologue of mammalian PIG-P, is a subunit of the initial enzyme for glycosylphosphatidylinositol anchor biosynthesis.Eukaryot Cell. 2005 Nov;4(11):1801-7. doi: 10.1128/EC.4.11.1801-1807.2005. Eukaryot Cell. 2005. PMID: 16278447 Free PMC article.
References
-
- Broach, J. R. 1991. RAS genes in Saccharomyces cerevisiae: signal transduction in search of a pathway. Trends Genet. 7: 28–32. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

