Discontinuous and non-discontinuous subgenomic RNA transcription in a nidovirus
- PMID: 12456663
- PMCID: PMC136939
- DOI: 10.1093/emboj/cdf635
Discontinuous and non-discontinuous subgenomic RNA transcription in a nidovirus
Abstract
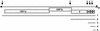
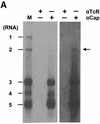
Arteri-, corona-, toro- and roniviruses are evolutionarily related positive-strand RNA viruses, united in the order Nidovirales. The best studied nidoviruses, the corona- and arteriviruses, employ a unique transcription mechanism, which involves discontinuous RNA synthesis, a process resembling similarity-assisted copy-choice RNA recombination. During infection, multiple subgenomic (sg) mRNAs are transcribed from a mirror set of sg negative-strand RNA templates. The sg mRNAs all possess a short 5' common leader sequence, derived from the 5' end of the genomic RNA. The joining of the non-contiguous 'leader' and 'body' sequences presumably occurs during minus-strand synthesis. To study whether toroviruses use a similar transcription mechanism, we characterized the 5' termini of the genome and the four sg mRNAs of Berne virus (BEV). We show that BEV mRNAs 3-5 lack a leader sequence. Surprisingly, however, RNA 2 does contain a leader, identical to the 5'-terminal 18 residues of the genome. Apparently, BEV combines discontinuous and non-discontinuous RNA synthesis to produce its sg mRNAs. Our findings have important implications for the understanding of the mechanism and evolution of nidovirus transcription.
Figures


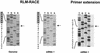





Similar articles
-
Torovirus non-discontinuous transcription: mutational analysis of a subgenomic mRNA promoter.J Virol. 2005 Jul;79(13):8275-81. doi: 10.1128/JVI.79.13.8275-8281.2005. J Virol. 2005. PMID: 15956573 Free PMC article.
-
Nidovirus transcription: how to make sense...?J Gen Virol. 2006 Jun;87(Pt 6):1403-1421. doi: 10.1099/vir.0.81611-0. J Gen Virol. 2006. PMID: 16690906 Review.
-
Discontinuous subgenomic RNA synthesis in arteriviruses is guided by an RNA hairpin structure located in the genomic leader region.J Virol. 2005 May;79(10):6312-24. doi: 10.1128/JVI.79.10.6312-6324.2005. J Virol. 2005. PMID: 15858015 Free PMC article.
-
Arterivirus discontinuous mRNA transcription is guided by base pairing between sense and antisense transcription-regulating sequences.Proc Natl Acad Sci U S A. 1999 Oct 12;96(21):12056-61. doi: 10.1073/pnas.96.21.12056. Proc Natl Acad Sci U S A. 1999. PMID: 10518575 Free PMC article.
-
A new model for coronavirus transcription.Adv Exp Med Biol. 1998;440:215-9. doi: 10.1007/978-1-4615-5331-1_26. Adv Exp Med Biol. 1998. PMID: 9782283 Review.
Cited by
-
Effects of modification of the transcription initiation site context on citrus tristeza virus subgenomic RNA synthesis.J Virol. 2003 Sep;77(17):9232-43. doi: 10.1128/jvi.77.17.9232-9243.2003. J Virol. 2003. PMID: 12915539 Free PMC article.
-
Crystal structure and functional analysis of the SARS-coronavirus RNA cap 2'-O-methyltransferase nsp10/nsp16 complex.PLoS Pathog. 2011 May;7(5):e1002059. doi: 10.1371/journal.ppat.1002059. Epub 2011 May 26. PLoS Pathog. 2011. PMID: 21637813 Free PMC article.
-
Structure-function analysis of the nsp14 N7-guanine methyltransferase reveals an essential role in Betacoronavirus replication.Proc Natl Acad Sci U S A. 2021 Dec 7;118(49):e2108709118. doi: 10.1073/pnas.2108709118. Proc Natl Acad Sci U S A. 2021. PMID: 34845015 Free PMC article.
-
Activation of the autophagy pathway by Torovirus infection is irrelevant for virus replication.PLoS One. 2019 Jul 15;14(7):e0219428. doi: 10.1371/journal.pone.0219428. eCollection 2019. PLoS One. 2019. PMID: 31306441 Free PMC article.
-
Torovirus non-discontinuous transcription: mutational analysis of a subgenomic mRNA promoter.J Virol. 2005 Jul;79(13):8275-81. doi: 10.1128/JVI.79.13.8275-8281.2005. J Virol. 2005. PMID: 15956573 Free PMC article.
References
-
- Bochnig P., Reuter,R., Bringmann,P. and Luhrmann,R. (1987) A monoclonal antibody against 2,2,7-trimethylguanosine that reacts with intact, class U, small nuclear ribonucleoproteins as well as with 7-methylguanosine-capped RNAs. Eur. J. Biochem., 168, 461–467. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

