The 3' untranslated region of human vimentin mRNA interacts with protein complexes containing eEF-1gamma and HAX-1
- PMID: 12466525
- PMCID: PMC137969
- DOI: 10.1093/nar/gkf656
The 3' untranslated region of human vimentin mRNA interacts with protein complexes containing eEF-1gamma and HAX-1
Abstract
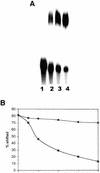
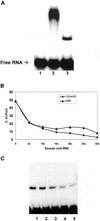
Previously, we have shown that the vimentin 3' untranslated region (3'UTR) contains a highly conserved region, which is sufficient for the perinuclear localization of a reporter mRNA. This region was shown to specifically bind protein(s) by band shift analyses. UV-cross-linking studies suggest these proteins are 46- and 35-kDa in mass. Here, we have used this sequence as 'bait' to isolate RNA binding proteins using the yeast three-hybrid method. This technique relies on a functional assay detecting bona fide RNA-protein interaction in vivo. Three cDNA isolates, HAX-1, eEF-1gamma and hRIP, code for proteins of a size consistent with in vitro cross- linking studies. In all cases, recombinant proteins were capable of binding RNA in vitro. Although hRIP is thought to be a general mRNA binding protein, this represents an unreported activity for eEF-1gamma and HAX-1. Moreover, HAX-1 binding appears to be specific to vimentin's 3'UTR. Both in vivo synthesized eEF-1gamma and HAX-1 proteins were 'pulled out' of HeLa whole cell extracts by binding to a RNA affinity column comprised of vimentin's 3'UTR. Moreover, size-fractionation of extracts results in the separation of large complexes containing either eEF-1gamma or HAX-1. Thus, in addition to their known functions, both eEF-1gamma and HAX-1 are RNA binding proteins, which suggests new roles in mRNA translation and/or perinuclear localization.
Figures






References
-
- Ross J. (1996) Control of mRNA stability in higher eukaryotes. Trends Genet., 12, 171–175. - PubMed
-
- Hunt T. (1989) On the translational control of suicide in red cell development. Trends Biochem. Sci., 14, 393–394. - PubMed
-
- Ostareck-Lederer A., Ostareck,D.H. and Hentze,M.W. (1998) Cytoplasmic regulatory functions of the KH-domain proteins hnRNPs K and E1/E2. Trends Biochem. Sci., 23, 409–411. - PubMed
-
- Wreden C., Verrotti,A.C., Schisa,J.A., Lieberfarb,M.E. and Strickland,S. (1997) Nanos and pumilio establish embryonic polarity in Drosophila by promoting posterior deadenylation of hunchback mRNA. Development, 124, 3015–3023. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

