Methylation of Xenopus CIRP2 regulates its arginine- and glycine-rich region-mediated nucleocytoplasmic distribution
- PMID: 12466543
- PMCID: PMC137953
- DOI: 10.1093/nar/gkf638
Methylation of Xenopus CIRP2 regulates its arginine- and glycine-rich region-mediated nucleocytoplasmic distribution
Abstract
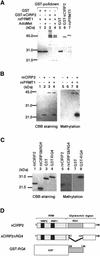
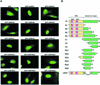
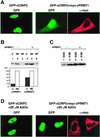
Cold-inducible RNA-binding protein (CIRP) was originally found in mammalian cells as a protein that is overexpressed upon a temperature downshift. Recently, we identified a Xenopus homolog of CIRP, termed xCIRP2, as a major cytoplasmic RNA-binding protein in oocytes. In this study we found by yeast two-hybrid screening that the Xenopus homolog of protein arginine N-methyltransferase 1 (xPRMT1) interacted with xCIRP2. We found that an arginine- and glycine-rich region of xCIRP2, termed the RG4 domain, was a target of xPRMT1 for methylation in vitro. xCIRP2 expressed in cultured cells accumulated in the nucleus as does mammalian CIRP. Interestingly, the RG4 domain was necessary for nuclear localization of xCIRP2. RG4-mediated nuclear accumulation of xCIRP2 was diminished in the presence of transcription inhibitors, suggesting that nuclear localization of xCIRP2 was dependent on ongoing transcription with RNA polymerase II. Analysis of interspecies heterokaryons revealed that xCIRP2 was capable of nucleocytoplasmic shuttling and the RG4 domain functioned as a nucleocytoplasmic shuttling signal. Methylation by overexpressed xPRMT1 caused cytoplasmic accumulation of xCIRP2. Possible implications of the relationship between regulation of intracellular localization and multiple functions of xCIRP2 will be discussed.
Figures



References
-
- Moore M.J. (2002) Nuclear RNA turnover. Cell, 108, 431–434. - PubMed
-
- Reed R. and Hurt,E. (2002) A conserved mRNA export machinery coupled to pre-mRNA splicing. Cell, 108, 523–531. - PubMed
-
- Shyu A.B. and Wilkinson,M.F. (2000) The double lives of shuttling mRNA binding proteins. Cell, 102, 135–138. - PubMed
-
- Siomi H. and Dreyfuss,G. (1997) RNA-binding proteins as regulators of gene expression. Curr. Opin. Genet. Dev., 7, 345–353. - PubMed
-
- Stutz F. and Rosbash,M. (1998) Nuclear RNA export. Genes Dev., 12, 3303–3319. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

