Discovery of RNA structural elements using evolutionary computation
- PMID: 12466557
- PMCID: PMC137967
- DOI: 10.1093/nar/gkf653
Discovery of RNA structural elements using evolutionary computation
Abstract
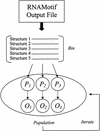
RNA molecules fold into characteristic secondary and tertiary structures that account for their diverse functional activities. Many of these RNA structures, or certain structural motifs within them, are thought to recur in multiple genes within a single organism or across the same gene in several organisms and provide a common regulatory mechanism. Search algorithms, such as RNAMotif, can be used to mine nucleotide sequence databases for these repeating motifs. RNAMotif allows users to capture essential features of known structures in detailed descriptors and can be used to identify, with high specificity, other similar motifs within the nucleotide database. However, when the descriptor constraints are relaxed to provide more flexibility, or when there is very little a priori information about hypothesized RNA structures, the number of motif 'hits' may become very large. Exhaustive methods to search for similar RNA structures over these large search spaces are likely to be computationally intractable. Here we describe a powerful new algorithm based on evolutionary computation to solve this problem. A series of experiments using ferritin IRE and SRP RNA stem-loop motifs were used to verify the method. We demonstrate that even when searching extremely large search spaces, of the order of 10(23) potential solutions, we could find the correct solution in a fraction of the time it would have taken for exhaustive comparisons.
Figures
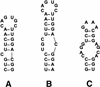


Similar articles
-
RNAMotif, an RNA secondary structure definition and search algorithm.Nucleic Acids Res. 2001 Nov 15;29(22):4724-35. doi: 10.1093/nar/29.22.4724. Nucleic Acids Res. 2001. PMID: 11713323 Free PMC article.
-
A method for aligning RNA secondary structures and its application to RNA motif detection.BMC Bioinformatics. 2005 Apr 7;6:89. doi: 10.1186/1471-2105-6-89. BMC Bioinformatics. 2005. PMID: 15817128 Free PMC article.
-
Finding a common motif of RNA sequences using genetic programming: the GeRNAMo system.IEEE/ACM Trans Comput Biol Bioinform. 2007 Oct-Dec;4(4):596-610. doi: 10.1109/tcbb.2007.1045. IEEE/ACM Trans Comput Biol Bioinform. 2007. PMID: 17975271
-
Energy-based RNA consensus secondary structure prediction in multiple sequence alignments.Methods Mol Biol. 2014;1097:125-41. doi: 10.1007/978-1-62703-709-9_7. Methods Mol Biol. 2014. PMID: 24639158 Review.
-
Graph Theoretical Methods and Workflows for Searching and Annotation of RNA Tertiary Base Motifs and Substructures.Int J Mol Sci. 2021 Aug 9;22(16):8553. doi: 10.3390/ijms22168553. Int J Mol Sci. 2021. PMID: 34445259 Free PMC article. Review.
Cited by
-
Number variation of high stability regions is correlated with gene functions.Genome Biol Evol. 2013;5(3):484-93. doi: 10.1093/gbe/evt020. Genome Biol Evol. 2013. PMID: 23407773 Free PMC article.
-
RNAProfile: an algorithm for finding conserved secondary structure motifs in unaligned RNA sequences.Nucleic Acids Res. 2004 Jun 15;32(10):3258-69. doi: 10.1093/nar/gkh650. Print 2004. Nucleic Acids Res. 2004. PMID: 15199174 Free PMC article.
-
Genome-wide bioinformatic prediction and experimental evaluation of potential RNA thermometers.Mol Genet Genomics. 2007 Nov;278(5):555-64. doi: 10.1007/s00438-007-0272-7. Epub 2007 Jul 24. Mol Genet Genomics. 2007. PMID: 17647020
-
Computational challenges, tools, and resources for analyzing co- and post-transcriptional events in high throughput.Wiley Interdiscip Rev RNA. 2015 May-Jun;6(3):291-310. doi: 10.1002/wrna.1274. Epub 2014 Dec 16. Wiley Interdiscip Rev RNA. 2015. PMID: 25515586 Free PMC article. Review.
-
Searching for IRES.RNA. 2006 Oct;12(10):1755-85. doi: 10.1261/rna.157806. Epub 2006 Sep 6. RNA. 2006. PMID: 16957278 Free PMC article. Review.
References
-
- Theil E.C. (1998) The iron responsive element (IRE) family of mRNA regulators. Regulation of iron transport and uptake compared in animals, plants and microorganisms. Met. Ions Biol. Syst., 35, 403–434. - PubMed
-
- Kim H.Y., LaVaute,T., Iwai,K., Klausner,R.D. and Rouault,T.A. (1996) Identification of a conserved and functional iron-responsive element in the 5′-untranslated region of mammalian mitochondrial aconitase. J. Biol. Chem., 271, 24226–24230. - PubMed
-
- Son S.Y. (1993) The structure and regulation of histone genes. Saenghwahak Nyusu, 13, 64–70.
-
- Le S.Y., Siddiqui,A. and Maizel,J.V.,Jr (1996) A common structural core in the internal ribosome entry sites of picornavirus, hepatitis C virus and pestivirus. Virus Genes, 12, 135–147. - PubMed

