Discovery of sulfated metabolites in mycobacteria with a genetic and mass spectrometric approach
- PMID: 12482950
- PMCID: PMC139265
- DOI: 10.1073/pnas.252514899
Discovery of sulfated metabolites in mycobacteria with a genetic and mass spectrometric approach
Abstract
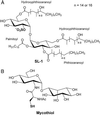
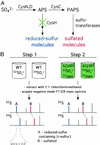
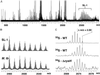
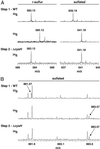
The study of the metabolome presents numerous challenges, first among them being the cataloging of its constituents. A step in this direction will be the development of tools to identify metabolites that share common structural features. The importance of sulfated molecules in cell-cell communication motivated us to develop a rapid two-step method for identifying these metabolites in microorganisms, particularly in pathogenic mycobacteria. Sulfurcontaining molecules were initially identified by mass spectral analysis of cell extracts from bacteria labeled metabolically with a stable sulfur isotope (34SO 4 2-). To differentiate sulfated from reduced-sulfur-containing molecules, we employed a mutant lacking the reductive branch of the sulfate assimilation pathway. In these sulfur auxotrophs, heavy sulfate is channeled exclusively into sulfated metabolites. The method was applied to the discovery of several new sulfated molecules in Mycobacterium tuberculosis and Mycobacterium smegmatis. Because a sulfur auxotrophic strain is the only requirement of the approach, many microorganisms can be studied in this manner. Such genetic engineering in combination with stable isotopic labeling can be applied to various metabolic pathways and their products.
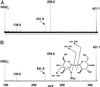
Figures
Similar articles
-
The metabolism of nitrosothiols in the Mycobacteria: identification and characterization of S-nitrosomycothiol reductase.Biochem J. 2003 Sep 15;374(Pt 3):657-66. doi: 10.1042/BJ20030642. Biochem J. 2003. PMID: 12809551 Free PMC article.
-
Mycothiol-dependent mycobacterial response to oxidative stress.FEBS Lett. 2006 May 15;580(11):2712-6. doi: 10.1016/j.febslet.2006.04.026. Epub 2006 Apr 21. FEBS Lett. 2006. PMID: 16643903
-
Mycothiol-deficient Mycobacterium smegmatis mutants are hypersensitive to alkylating agents, free radicals, and antibiotics.Antimicrob Agents Chemother. 2002 Nov;46(11):3348-55. doi: 10.1128/AAC.46.11.3348-3355.2002. Antimicrob Agents Chemother. 2002. PMID: 12384335 Free PMC article.
-
Mycothiol biochemistry.Arch Microbiol. 2002 Dec;178(6):388-94. doi: 10.1007/s00203-002-0469-4. Epub 2002 Sep 3. Arch Microbiol. 2002. PMID: 12420157 Review.
-
Drug targets in mycobacterial sulfur metabolism.Infect Disord Drug Targets. 2007 Jun;7(2):140-58. doi: 10.2174/187152607781001772. Infect Disord Drug Targets. 2007. PMID: 17970225 Free PMC article. Review.
Cited by
-
Structural characterization of a novel sulfated menaquinone produced by stf3 from Mycobacterium tuberculosis.ACS Chem Biol. 2008 Oct 17;3(10):619-24. doi: 10.1021/cb800145r. ACS Chem Biol. 2008. PMID: 18928249 Free PMC article.
-
New targets and inhibitors of mycobacterial sulfur metabolism.Infect Disord Drug Targets. 2013 Apr;13(2):85-115. doi: 10.2174/18715265113139990022. Infect Disord Drug Targets. 2013. PMID: 23808874 Free PMC article. Review.
-
Biosynthesis and Regulation of Sulfomenaquinone, a Metabolite Associated with Virulence in Mycobacterium tuberculosis.ACS Infect Dis. 2016 Nov 11;2(11):800-806. doi: 10.1021/acsinfecdis.6b00106. Epub 2016 Aug 15. ACS Infect Dis. 2016. PMID: 27933784 Free PMC article.
-
Bioaerosol mass spectrometry for rapid detection of individual airborne Mycobacterium tuberculosis H37Ra particles.Appl Environ Microbiol. 2005 Oct;71(10):6086-95. doi: 10.1128/AEM.71.10.6086-6095.2005. Appl Environ Microbiol. 2005. PMID: 16204525 Free PMC article.
-
Target Identification in Anti-Tuberculosis Drug Discovery.Int J Mol Sci. 2023 Jun 22;24(13):10482. doi: 10.3390/ijms241310482. Int J Mol Sci. 2023. PMID: 37445660 Free PMC article. Review.
References
-
- Kehoe J. W. & Bertozzi, C. R. (2000) Chem. Biol. 7, R57-R61. - PubMed
-
- Bowman K. G. & Bertozzi, C. R. (1999) Chem. Biol. 6, R9-R22. - PubMed
-
- Hemmerich S. & Rosen, S. D. (2000) Glycobiology 10, 849-856. - PubMed
-
- Roche P., Debelle, F., Maillet, F., Lerouge, P., Faucher, C., Truchet, G., Denarie, J. & Prome, J. C. (1991) Cell 67, 1131-1143. - PubMed
-
- Hanin M., Jabbouri, S., Quesada-Vincens, D., Freiberg, C., Perret, X., Prome, J. C., Broughton, W. J. & Fellay, R. (1997) Mol. Microbiol. 24, 1119-1129. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

