Human MI-ER1 alpha and beta function as transcriptional repressors by recruitment of histone deacetylase 1 to their conserved ELM2 domain
- PMID: 12482978
- PMCID: PMC140656
- DOI: 10.1128/MCB.23.1.250-258.2003
Human MI-ER1 alpha and beta function as transcriptional repressors by recruitment of histone deacetylase 1 to their conserved ELM2 domain
Abstract
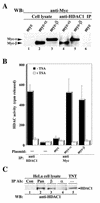
mi-er1 (previously called er1) was first isolated from Xenopus laevis embryonic cells as a novel fibroblast growth factor-regulated immediate-early gene. Xmi-er1 was shown to encode a nuclear protein with an N-terminal acidic transcription activation domain. The human orthologue of mi-er1 (hmi-er1) displays 91% similarity to the Xenopus sequence at the amino acid level and was shown to be upregulated in breast carcinoma cell lines and tumors. Alternative splicing at the 3' end of hmi-er1 produces two major isoforms, hMI-ER1alpha and hMI-ER1beta, which contain distinct C-terminal domains. In this study, we investigated the role of hMI-ER1alpha and hMI-ER1beta in the regulation of transcription. Using fusion proteins of hMI-ER1alpha or hMI-ER1beta tethered to the GAL4 DNA binding domain, we show that both isoforms, when recruited to the G5tkCAT minimal promoter, function to repress transcription. We demonstrate that this repressor activity is due to interaction and recruitment of a trichostatin A-sensitive histone deacetylase 1 (HDAC1). Furthermore, deletion analysis revealed that recruitment of HDAC1 to hMI-ER1alpha and hMI-ER1beta occurs through their common ELM2 domain. The ELM2 domain was first described in the Caenorhabditis elegans Egl-27 protein and is present in a number of SANT domain-containing transcription factors. This is the first report of a function for the ELM2 domain, highlighting its role in the regulation of transcription.
Figures
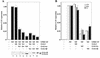
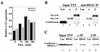



References
-
- Aasland, R., A. F. Stewart, and T. Gibson. 1996. The SANT domain: a putative DNA-binding domain in the SWI-SNF and ADA complexes, the transcriptional co-repressor N-CoR and TFIIIB. Trends Biochem. Sci. 21:87-88. - PubMed
-
- Aranda, A., and A. Pascual. 2001. Nuclear hormone receptors and gene expression. Physiol. Rev. 81:1270-1304. - PubMed
-
- Ayer, D. E. 1999. Histone deacetylases: transcriptional repression with SINers and NuRDs. Trends Cell Biol. 9:193-198. - PubMed
-
- Brubaker, K., S. M. Cowley, K. Huang, L. Loo, G. S. Yochum, D. E. Ayer, R. N. Eisenman, and I. Radhakrishnan. 2000. Solution structure of the interacting domains of the Mad-Sin3 complex: implications for recruitment of a chromatin-modifying complex. Cell 103:655-665. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
