A novel domain within the DEAD-box protein DP103 is essential for transcriptional repression and helicase activity
- PMID: 12482992
- PMCID: PMC140651
- DOI: 10.1128/MCB.23.1.414-423.2003
A novel domain within the DEAD-box protein DP103 is essential for transcriptional repression and helicase activity
Abstract
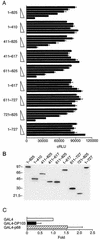
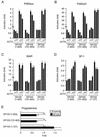
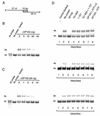
Members of the DEAD-box family of helicases, distinguished by a core characteristic sequence of Asp-Glu-Ala-Asp, are expressed in a wide range of prokaryotes and eukaryotes and exhibit diverse cellular functions, including DNA transcription, recombination and repair, RNA processing, translation, and posttranslational regulation. Although ubiquitous, the function of most DEAD-box proteins is unknown. We and others have recently cloned DP103, which harbors conserved DEAD-box, helicase, and ATPase domains in its N terminus. DP103 (also termed Gemin3 and DDX20) interacts with SF-1, SMN, EBNA2, and EBNA3C in mammalian cells. Here we demonstrate that a discrete domain within the nonconserved C-terminal region of DP103 directly interacts with SF-1. This domain exhibits an autonomous repression function and is necessary and sufficient for repressing the transcriptional activity of SF-1. Furthermore, intact DP103 exhibits helicase activity. Importantly, the C-terminal domain is obligatory but not sufficient for this unwinding activity of DP103. Together, our results support a novel paradigm for transcriptional repression and demonstrate the bifunctional role of the C-terminal domain of DP103.
Figures

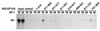

Similar articles
-
The DEAD box protein DP103 is a regulator of steroidogenic factor-1.Mol Endocrinol. 2001 Jan;15(1):69-79. doi: 10.1210/mend.15.1.0580. Mol Endocrinol. 2001. PMID: 11145740
-
The DEAD-box protein DP103 (Ddx20 or Gemin-3) represses orphan nuclear receptor activity via SUMO modification.Mol Cell Biol. 2005 Mar;25(5):1879-90. doi: 10.1128/MCB.25.5.1879-1890.2005. Mol Cell Biol. 2005. PMID: 15713642 Free PMC article.
-
The Ddx20/DP103 dead box protein represses transcriptional activation by Egr2/Krox-20.J Biol Chem. 2004 Mar 5;279(10):9056-63. doi: 10.1074/jbc.M309308200. Epub 2003 Dec 29. J Biol Chem. 2004. PMID: 14699164
-
The multiple lives of DEAD-box RNA helicase DP103/DDX20/Gemin3.Biochem Soc Trans. 2018 Apr 17;46(2):329-341. doi: 10.1042/BST20180016. Epub 2018 Mar 9. Biochem Soc Trans. 2018. PMID: 29523774 Review.
-
Modulation of transcriptional activity of the DEAD-box family of RNA helicases, p68 (Ddx5) and DP103 (Ddx20), by SUMO modification.Biochem Soc Trans. 2007 Dec;35(Pt 6):1427-9. doi: 10.1042/BST0351427. Biochem Soc Trans. 2007. PMID: 18031238 Review.
Cited by
-
Impact of EBV essential nuclear protein EBNA-3C on B-cell proliferation and apoptosis.Future Microbiol. 2013 Mar;8(3):323-52. doi: 10.2217/fmb.12.147. Future Microbiol. 2013. PMID: 23464371 Free PMC article. Review.
-
Recognition of two distinct elements in the RNA substrate by the RNA-binding domain of the T. thermophilus DEAD box helicase Hera.Nucleic Acids Res. 2013 Jul;41(12):6259-72. doi: 10.1093/nar/gkt323. Epub 2013 Apr 25. Nucleic Acids Res. 2013. PMID: 23625962 Free PMC article.
-
Conserved requirement for DEAD-box RNA helicase Gemin3 in Drosophila oogenesis.BMC Res Notes. 2012 Feb 23;5:120. doi: 10.1186/1756-0500-5-120. BMC Res Notes. 2012. PMID: 22361416 Free PMC article.
-
The DEAD-box helicase eIF4A: paradigm or the odd one out?RNA Biol. 2013 Jan;10(1):19-32. doi: 10.4161/rna.21966. Epub 2012 Sep 20. RNA Biol. 2013. PMID: 22995829 Free PMC article. Review.
-
Human placental microRNAs and preeclampsia.Biol Reprod. 2013 May 23;88(5):130. doi: 10.1095/biolreprod.113.107805. Print 2013 May. Biol Reprod. 2013. PMID: 23575145 Free PMC article. Review.
References
-
- Campbell, L., K. M. Hunter, P. Mohaghegh, J. M. Tinsley, M. A. Brasch, and K. E. Davies. 2000. Direct interaction of Smn with dp103, a putative RNA helicase: a role for Smn in transcription regulation? Hum. Mol. Genet. 9:1093-1100. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
