Protein hydrogen exchange mechanism: local fluctuations
- PMID: 12493838
- PMCID: PMC2312409
- DOI: 10.1110/ps.0225803
Protein hydrogen exchange mechanism: local fluctuations
Abstract
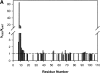
Experiments were done to study the dynamic structural motions that determine protein hydrogen exchange (HX) behavior. The replacement of a solvent-exposed lysine residue with glycine (Lys8Gly) in a helix of recombinant cytochrome c does not perturb the native structure, but it entropically potentiates main-chain flexibility and thus can promote local distortional motions and large-scale unfolding. The mutation accelerates amide hydrogen exchange of the mutated residue by about 50-fold, neighboring residues in the same helix by less, and residues elsewhere in the protein not at all, except for Leu98, which registers the change in global stability. The pattern of HX changes shows that the coupled structural distortions that dominate exchange can be several residues in extent, but they expose to exchange only one amide NH at a time. This "local fluctuation" mode of hydrogen exchange may be generally recognized by disparate near-neighbor rates and a low dependence on destabilants (denaturant, temperature, pressure). In contrast, concerted unfolding reactions expose multiple neighboring amide NHs with very similar computed protection factors, and they show marked destabilant sensitivity. In both modes, ionic hydrogen exchange catalysts attack from the bulk solvent without diffusing through the protein matrix.
Figures



Similar articles
-
Determinants of protein hydrogen exchange studied in equine cytochrome c.Protein Sci. 1998 Mar;7(3):739-45. doi: 10.1002/pro.5560070323. Protein Sci. 1998. PMID: 9541406 Free PMC article.
-
Conformational dynamics of cytochrome c: correlation to hydrogen exchange.Proteins. 1999 Aug 1;36(2):175-91. Proteins. 1999. PMID: 10398365
-
The asparagine-stabilized beta-turn of apamin: contribution to structural stability from dynamics simulation and amide hydrogen exchange analysis.Biochemistry. 2000 Dec 26;39(51):15944-52. doi: 10.1021/bi002044q. Biochemistry. 2000. PMID: 11123921
-
Structural description of acid-denatured cytochrome c by hydrogen exchange and 2D NMR.Biochemistry. 1990 Nov 20;29(46):10433-7. doi: 10.1021/bi00498a001. Biochemistry. 1990. PMID: 2176867 Review.
-
Protein analysis by hydrogen exchange mass spectrometry.Annu Rev Biophys Biomol Struct. 2003;32:1-25. doi: 10.1146/annurev.biophys.32.110601.142417. Epub 2003 Feb 18. Annu Rev Biophys Biomol Struct. 2003. PMID: 12598366 Review.
Cited by
-
Characterization of H/D exchange in type 1 pili by proton-detected solid-state NMR and molecular dynamics simulations.J Biomol NMR. 2019 Jul;73(6-7):281-291. doi: 10.1007/s10858-019-00247-3. Epub 2019 Apr 26. J Biomol NMR. 2019. PMID: 31028572 Free PMC article.
-
NumSimEX: A method using EXX hydrogen exchange mass spectrometry to map the energetics of protein folding landscapes.Protein Sci. 2025 Feb;34(2):e70045. doi: 10.1002/pro.70045. Protein Sci. 2025. PMID: 39865386 Free PMC article.
-
Differential accessibilities of dibasic prohormone processing sites of proenkephalin to the aqueous environment revealed by H-D exchange mass spectrometry.Biochemistry. 2009 Feb 24;48(7):1604-12. doi: 10.1021/bi801888j. Biochemistry. 2009. PMID: 19173595 Free PMC article.
-
Role of partial protein unfolding in alcohol-induced protein aggregation.Proteins. 2010 Sep;78(12):2625-37. doi: 10.1002/prot.22778. Proteins. 2010. PMID: 20597088 Free PMC article.
-
A protein folding pathway with multiple folding intermediates at atomic resolution.Proc Natl Acad Sci U S A. 2005 Apr 5;102(14):5026-31. doi: 10.1073/pnas.0501372102. Epub 2005 Mar 25. Proc Natl Acad Sci U S A. 2005. PMID: 15793003 Free PMC article.
References
-
- ———. 1994. Protein stability parameters measured by hydrogen exchange. Proteins Struct. Funct. Genet. 20 4–14. - PubMed
-
- Barksdale, A.D. and Rosenberg, A. 1982. Acquisition and interpretation of hydrogen exchange data from peptides, polymers, and proteins. In Methods of biochemical analysis (ed. D. Glick), p. 113. John Wiley & Sons, New York, NY. - PubMed
-
- Berger, A. and Linderstrøm-Lang, K. 1957. Deuterium exchange of poly-D,L-alanine in aqueous solution. Arch. Biochem. Biophys. 69 106–118. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

