Biological functions of the ISWI chromatin remodeling complex NURF
- PMID: 12502740
- PMCID: PMC187504
- DOI: 10.1101/gad.1032202
Biological functions of the ISWI chromatin remodeling complex NURF
Abstract
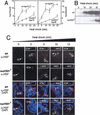
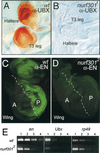
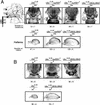
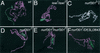
The nucleosome remodeling factor (NURF) is one of several ISWI-containing protein complexes that catalyze ATP-dependent nucleosome sliding and facilitate transcription of chromatin in vitro. To establish the physiological requirements of NURF, and to distinguish NURF genetically from other ISWI-containing complexes, we isolated mutations in the gene encoding the large NURF subunit, nurf301. We confirm that NURF is required for transcription activation in vivo. In animals lacking NURF301, heat-shock transcription factor binding to and transcription of the hsp70 and hsp26 genes are impaired. Additionally, we show that NURF is required for homeotic gene expression. Consistent with this, nurf301 mutants recapitulate the phenotypes of Enhancer of bithorax, a positive regulator of the Bithorax-Complex previously localized to the same genetic interval. Finally, mutants in NURF subunits exhibit neoplastic transformation of larval blood cells that causes melanotic tumors to form.
Figures



References
-
- Andres AJ, Thummel CS. Methods for quantitative analysis of transcription in larvae and prepupae. Methods Cell Biol. 1994;44:565–573. - PubMed
-
- Bender W, Akam ME, Karch F, Beachy PA, Peifer M, Spierer P, Lewis EB, Hogness DS. Molecular genetics of the Bithorax Complex in Drosophila melanogaster. Science. 1983;221:23–29. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
