Elevated phospholipase D activity in H-Ras- but not K-Ras-transformed cells by the synergistic action of RalA and ARF6
- PMID: 12509462
- PMCID: PMC151535
- DOI: 10.1128/MCB.23.2.645-654.2003
Elevated phospholipase D activity in H-Ras- but not K-Ras-transformed cells by the synergistic action of RalA and ARF6
Abstract
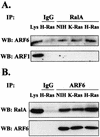
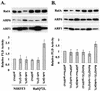
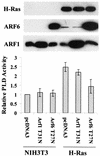
Phospholipase D (PLD) activity is elevated in response to the oncogenic stimulus of H-Ras but not K-Ras. H-Ras and K-Ras have been reported to localize to different membrane microdomains, with H-Ras localizing to caveolin-enriched light membrane fractions. We reported previously that PLD activity elevated in response to mitogenic stimulation is restricted to the caveolin-enriched light membrane fractions. PLD activity in H-Ras-transformed cells is dependent upon RalA, and consistent with a lack of elevated PLD activity in K-Ras-transformed cells, RalA was not activated in K-Ras-transformed cells. Although H-Ras-induced PLD activity is dependent upon RalA, an activated mutant of RalA is not sufficient to elevate PLD activity. We reported previously that RalA interacts with PLD activating ADP ribosylation factor (ARF) proteins. In cells transformed by H-Ras, we found increased coprecipitation of ARF6 with RalA. Moreover, ARF6 colocalized with RalA in light membrane fractions. Interestingly, ARF6 protein levels were elevated in H-Ras- but not K-Ras-transformed cells. A dominant-negative mutant of ARF6 inhibited PLD activity in H-Ras-transformed NIH 3T3 cells. Activated mutants of either ARF6 or RalA were not sufficient to elevate PLD activity in NIH 3T3 cells; however, expression of both activated RalA and activated ARF6 in NIH 3T3 cells led to increased PLD activity. These data suggest a model whereby H-Ras stimulates the activation of both RalA and ARF6, which together lead to the elevation of PLD activity.
Figures






Similar articles
-
Involvement of Ral GTPase in v-Src-induced phospholipase D activation.Nature. 1995 Nov 23;378(6555):409-12. doi: 10.1038/378409a0. Nature. 1995. PMID: 7477381
-
Phospholipase D and RalA cooperate with the epidermal growth factor receptor to transform 3Y1 rat fibroblasts.Mol Cell Biol. 2000 Jan;20(2):462-7. doi: 10.1128/MCB.20.2.462-467.2000. Mol Cell Biol. 2000. PMID: 10611224 Free PMC article.
-
Functional association between Arf and RalA in active phospholipase D complex.Proc Natl Acad Sci U S A. 1998 Mar 31;95(7):3632-7. doi: 10.1073/pnas.95.7.3632. Proc Natl Acad Sci U S A. 1998. PMID: 9520417 Free PMC article.
-
Phospholipase D stimulation by receptor tyrosine kinases mediated by protein kinase C and a Ras/Ral signaling cascade.J Biol Chem. 1999 Dec 3;274(49):34691-8. doi: 10.1074/jbc.274.49.34691. J Biol Chem. 1999. PMID: 10574935
-
The regulation of phospholipase D by inositol phospholipids and small GTPases.FEBS Lett. 2002 Oct 30;531(1):62-4. doi: 10.1016/s0014-5793(02)03410-5. FEBS Lett. 2002. PMID: 12401204 Review.
Cited by
-
Ral GTPases regulate neurite branching through GAP-43 and the exocyst complex.J Cell Biol. 2005 Dec 5;171(5):857-69. doi: 10.1083/jcb.200507061. J Cell Biol. 2005. PMID: 16330713 Free PMC article.
-
Heterogeneity of phosphatidic acid levels and distribution at the plasma membrane in living cells as visualized by a Föster resonance energy transfer (FRET) biosensor.J Biol Chem. 2010 Nov 12;285(46):35979-87. doi: 10.1074/jbc.M110.153007. Epub 2010 Sep 8. J Biol Chem. 2010. PMID: 20826779 Free PMC article.
-
PLD3 in Alzheimer's disease.Mol Neurobiol. 2015 Apr;51(2):480-6. doi: 10.1007/s12035-014-8779-5. Epub 2014 Jun 17. Mol Neurobiol. 2015. PMID: 24935720 Review.
-
ADP-Ribosylation Factor 1 Regulates Proliferation, Migration, and Fusion in Early Stage of Osteoclast Differentiation.Int J Mol Sci. 2015 Dec 9;16(12):29305-14. doi: 10.3390/ijms161226168. Int J Mol Sci. 2015. PMID: 26690137 Free PMC article.
-
RalA activation at nascent lamellipodia of epidermal growth factor-stimulated Cos7 cells and migrating Madin-Darby canine kidney cells.Mol Biol Cell. 2004 Jun;15(6):2549-57. doi: 10.1091/mbc.e03-11-0857. Epub 2004 Mar 19. Mol Biol Cell. 2004. PMID: 15034142 Free PMC article.
References
-
- Aguirre Ghiso, J., P. Frankel, Z. Lu, H. Jiang, E. Farias, A. Olsen, L. A. Feig, E. B. de Kier Joffe, and D. A. Foster. 1999. RalA requirement for v-Src- and v-Ras-induced tumorigenicity and overproduction of urokinase-type plasminogen activator and metalloproteases. Oncogene 18:4718-4725. - PubMed
-
- Alam, M. S., Y. Banno, S. Nakashima, and Y. Nozawa. 1995. Defective phospholipase D activation in Ki-ras-transformed NIH3T3 cells: evidence for downstream effector of PLC-γ1 in PDGF-mediated signal transduction. Biochem. Biophys. Res. Commun. 207:460-466. - PubMed
-
- Bos, J. L. 1989. Ras oncogenes in human cancer: a review. Cancer Res. 49:4682-4689. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
