LEAFY COTYLEDON1-LIKE defines a class of regulators essential for embryo development
- PMID: 12509518
- PMCID: PMC143447
- DOI: 10.1105/tpc.006973
LEAFY COTYLEDON1-LIKE defines a class of regulators essential for embryo development
Abstract
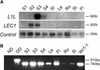
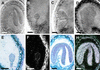
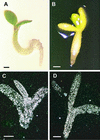
Arabidopsis LEAFY COTYLEDON1 (LEC1) is a critical regulator required for normal development during the early and late phases of embryogenesis that is sufficient to induce embryonic development in vegetative cells. LEC1 encodes a HAP3 subunit of the CCAAT binding transcription factor. We show that the 10 Arabidopsis HAP3 (AHAP3) subunits can be divided into two classes based on sequence identity in their central, conserved B domain. LEC1 and its most closely related subunit, LEC1-LIKE (L1L), constitute LEC1-type AHAP3 subunits, whereas the remaining AHAP3 subunits are designated non-LEC1-type. Similar to LEC1, L1L is expressed primarily during seed development. However, suppression of L1L gene expression induced defects in embryo development that differed from those of lec1 mutants, suggesting that LEC1 and L1L play unique roles in embryogenesis. We show that L1L expressed under the control of DNA sequences flanking the LEC1 gene suppressed genetically the lec1 mutation, suggesting that the LEC1-type B domains of L1L and LEC1 are critical for their function in embryogenesis. Our results also suggest that LEC1-type HAP3 subunits arose from a common origin uniquely in plants. Thus, L1L, an essential regulator of embryo development, defines a unique class of plant HAP3 subunits.
Figures




References
-
- Albani, D., and Robert, L.S. (1995). Cloning and characterization of a Brassica napus gene encoding a homologue of the B subunit of a heteromeric CCAAT-binding factor. Gene 167, 209–213. - PubMed
-
- Ausubel, F.M., Brent, R., Kingston, R.E., Moore, D.D., Seidman, J.G., and Smith, J.A. (1995). Short Protocols in Molecular Biology, 3rd ed. (New York: John Wiley & Sons).
-
- Bechtold, N., Ellis, J., and Pelletier, G. (1993). In planta Agrobacterium mediated gene transfer by infiltration of adult Arabidopsis thaliana plants. C. R. Acad. Sci. Paris 316, 1194–1199.
-
- Coustry, F., Maity, S.N., Sinha, S., and de Crombrugghe, B. (1996). The transcriptional activity of the CCAAT-binding factor CBF is mediated by two distinct activation domains, one in the CBF-B subunit and the other in the CBF-C subunit. J. Biol. Chem. 272, 14485–14491. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

