The Structure Superposition Database
- PMID: 12520064
- PMCID: PMC165574
- DOI: 10.1093/nar/gkg127
The Structure Superposition Database
Abstract
The need for new tools for investigating biological systems on a large scale is becoming acute, particularly with respect to computationally intensive analyses such as comparisons of many three-dimensional protein structures. Structure superposition is a valuable approach for understanding evolutionary relationships and for the prediction of function. But while available tools are adequate for generating and viewing superpositions of single pairs of protein structures, these tools are generally too cumbersome and time-consuming for examining multiple superpositions. To address this need, we have created the Structure Superposition Database (SSD) for accessing, viewing and understanding large sets of structure superposition data. The initial implementation of the SSD contains the results of pairwise, all-by-all superpositions of a representative set of 115 (beta/alpha)8 barrel structures (TIM barrels). Future plans call for extending the database to include representative structure superpositions for many additional folds. The SSD can be browsed with a user interface module developed as an extension to Chimera, an extensible molecular modeling program. Features of the user interface module facilitate viewing multiple superpositions together. The SSD interface module can be downloaded from http://ssd.rbvi.ucsf.edu.
Figures
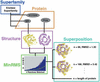


References
-
- Burley S.K., Almo,S.C., Bonanno,J.B., Capel,M., Chance,M.R., Gaasterland,T., Lin,D., Sali,A., Studier,F.W. and Swaminathan,S. (1999) Structural genomics: beyond the human genome project. Nature Genet., 23, 151–157. - PubMed
-
- Holm L. and Sander,C. (1996) Mapping the Protein Universe. Science, 273, 595–603. - PubMed
-
- Young M.M., Skillman,G. and Kuntz,I.D. (1999) A rapid method for exploring the protein structure universe. Proteins, 34, 317–332. - PubMed
-
- Dietmann S. and Holm,L. (2001) Identification of homology in protein structure classification. Nature Struct. Biol., 8, 953–957. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

