Expression and assembly of a fully active antibody in algae
- PMID: 12522151
- PMCID: PMC141013
- DOI: 10.1073/pnas.0237108100
Expression and assembly of a fully active antibody in algae
Abstract
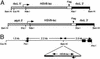
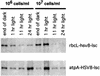
Although combinatorial antibody libraries have solved the problem of access to large immunological repertoires, efficient production of these complex molecules remains a problem. Here we demonstrate the efficient expression of a unique large single-chain (lsc) antibody in the chloroplast of the unicellular, green alga, Chlamydomonas reinhardtii. We achieved high levels of protein accumulation by synthesizing the lsc gene in chloroplast codon bias and by driving expression of the chimeric gene using either of two C. reinhardtii chloroplast promoters and 5' and 3' RNA elements. This lsc antibody, directed against glycoprotein D of the herpes simplex virus, is produced in a soluble form by the alga and assembles into higher order complexes in vivo. Aside from dimerization by disulfide bond formation, the antibody undergoes no detectable posttranslational modification. We further demonstrate that accumulation of the antibody can be modulated by the specific growth regime used to culture the alga, and by the choice of 5' and 3' elements used to drive expression of the antibody gene. These results demonstrate the utility of alga as an expression platform for recombinant proteins, and describe a new type of single chain antibody containing the entire heavy chain protein, including the Fc domain.
Figures




References
-
- Dove A. Nat Biotechnol. 2002;20:777–779. - PubMed
-
- Motmans K, Bouche B. Antibodies: The Next Generation. New York: Auerbach Grayson; 2000.
-
- Hiatt A, Cafferky R, Bowdish K. Nature. 1989;342:76–78. - PubMed
-
- Ma J, Lehner T, Stabila P, Fux C I, Hiatt A. Eur J Immunol. 1994;24:131–138. - PubMed
-
- Ma J, Hiatt A, Hein M, Vine N D, Wang F, Satabila P, van Dollweerd C, Mostov K, Lehner T. Science. 1995;268:716–719. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

