Optimized expression and specific activity of IL-12 by directed molecular evolution
- PMID: 12529500
- PMCID: PMC298744
- DOI: 10.1073/pnas.0237327100
Optimized expression and specific activity of IL-12 by directed molecular evolution
Abstract
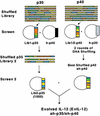
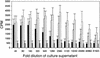
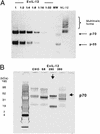
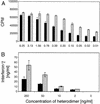
DNA delivery of IL-12 has shown promise in reducing the toxic side effects associated with administration of recombinant human (h)IL-12 protein while maintaining the ability to inhibit tumor growth and abolish tumor metastases in animal models. We have developed a more potent version of IL-12 by using DNA shuffling and screening to improve its expression in human cells and specific activity on human T cells. The most improved evolved IL-12 (EvIL-12) derived from seven mammalian genes encoding both the p35 and p40 subunits of IL-12 showed a 128-fold improvement in human T cell proliferation compared with native hIL-12 during the initial screening of supernatants from transected cells. When purified hIL-12 and EvIL-12 proteins were compared in vitro in human T cell proliferation and Th1 differentiation assays, it was demonstrated that EvIL-12 exhibited a concomitant 10-fold increase in the specific activity of the protein compared with hIL-12. Furthermore, DNA shuffling improved the level of expression and homogeneity of the heterodimer synthesized by 293 human embryonic kidney cells transfected with EvIL-12 by at least 10-fold. Molecular analysis of the variant revealed strategic placement of amino acid substitutions that potentially may facilitate heterodimer formation and product expression. The enhanced expression and biological activity of EvIL-12 may improve the effectiveness of IL-12 gene-based vaccines and therapeutics without the toxic side effects sometimes associated with hIL-12 protein administration.
Figures


Similar articles
-
[Expression of human interleukin-12 in Baculovirus expression system].Wei Sheng Wu Xue Bao. 2001 Feb;41(1):35-42. Wei Sheng Wu Xue Bao. 2001. PMID: 12549186 Chinese.
-
Construction of vectors expressing bioactive heterodimeric and single-chain murine interleukin-12 for gene therapy.Hum Gene Ther. 1998 Mar 1;9(4):457-65. doi: 10.1089/hum.1998.9.4-457. Hum Gene Ther. 1998. PMID: 9525307
-
Engineering N-glycosylation mutations in IL-12 enhances sustained cytotoxic T lymphocyte responses for DNA immunization.Nat Biotechnol. 2002 Apr;20(4):381-6. doi: 10.1038/nbt0402-381. Nat Biotechnol. 2002. PMID: 11923845
-
[Novel vaccines against M. tuberculosis].Kekkaku. 2006 Dec;81(12):745-51. Kekkaku. 2006. PMID: 17240920 Review. Japanese.
-
IL-12: the role of p40 versus p75.Scand J Immunol. 2002 Jul;56(1):1-11. doi: 10.1046/j.1365-3083.2002.01101.x. Scand J Immunol. 2002. PMID: 12100467 Review.
Cited by
-
The conversion of 3' UTRs into coding regions.Mol Biol Evol. 2007 Feb;24(2):457-64. doi: 10.1093/molbev/msl172. Epub 2006 Nov 10. Mol Biol Evol. 2007. PMID: 17099057 Free PMC article.
-
Revealing biases inherent in recombination protocols.BMC Biotechnol. 2007 Nov 14;7:77. doi: 10.1186/1472-6750-7-77. BMC Biotechnol. 2007. PMID: 18001472 Free PMC article.
-
A directed molecular evolution approach to improved immunogenicity of the HIV-1 envelope glycoprotein.PLoS One. 2011;6(6):e20927. doi: 10.1371/journal.pone.0020927. Epub 2011 Jun 29. PLoS One. 2011. PMID: 21738594 Free PMC article.
-
The rarity of gene shuffling in conserved genes.Genome Biol. 2005;6(6):R50. doi: 10.1186/gb-2005-6-6-r50. Epub 2005 May 9. Genome Biol. 2005. PMID: 15960802 Free PMC article.
-
Laboratory-directed protein evolution.Microbiol Mol Biol Rev. 2005 Sep;69(3):373-92. doi: 10.1128/MMBR.69.3.373-392.2005. Microbiol Mol Biol Rev. 2005. PMID: 16148303 Free PMC article. Review.
References
-
- Gately M K, Renzetti L M, Magram J, Stern A S, Adorin I L, Gubler U, Presky D H. Annu Rev Immunol. 1998;16:495–521. - PubMed
-
- Trinchieri G, Scott P. Curr Top Microbiol Immunol. 1999;238:57–78. - PubMed
-
- Greenberger M J, Kunkel S, Strieter R, Lukacs N, Bramson J, Gauldie J, Graham F, Hitt M, Danforth J, Standiford T. J Immunol. 1966;157:3006–3012. - PubMed
-
- Silva R, Pais T, Appelberg R. J Immunol. 1998;161:5578–5585. - PubMed
-
- Miller M, Skeen M, Ziegler H. Cell Immunol. 1998;184:92–104. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

