Erythroid differentiation sensitizes K562 leukemia cells to TRAIL-induced apoptosis by downregulation of c-FLIP
- PMID: 12556488
- PMCID: PMC141136
- DOI: 10.1128/MCB.23.4.1278-1291.2003
Erythroid differentiation sensitizes K562 leukemia cells to TRAIL-induced apoptosis by downregulation of c-FLIP
Abstract
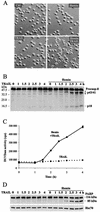
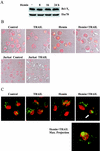
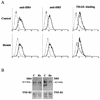
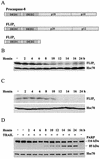
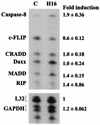
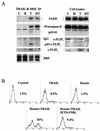
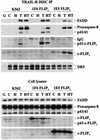
Regulation of the apoptotic threshold is of great importance in the homeostasis of both differentiating and fully developed organ systems. Triggering differentiation has been employed as a strategy to inhibit cell proliferation and accelerate apoptosis in malignant cells, in which the apoptotic threshold is often characteristically elevated. To better understand the mechanisms underlying differentiation-mediated regulation of apoptosis, we have studied death receptor responses during erythroid differentiation of K562 erythroleukemia cells, which normally are highly resistant to tumor necrosis factor (TNF) alpha-, FasL-, and TRAIL-induced apoptosis. However, upon hemin-mediated erythroid differentiation, K562 cells specifically lost their resistance to TNF-related apoptosis-inducing ligand (TRAIL), which efficiently killed the differentiating cells independently of mitochondrial apoptotic signaling. Concomitantly with the increased sensitivity, the expression of both c-FLIP splicing variants, c-FLIP(L) and c-FLIP(S), was downregulated, resulting in an altered caspase 8 recruitment and cleavage in the death-inducing signaling complex (DISC). Stable overexpression of both c-FLIP(L) and c-FLIP(S) rescued the cells from TRAIL-mediated apoptosis with isoform-specific effects on DISC-recruited caspase 8. Our results show that c-FLIP(L) and c-FLIP(S) potently control TRAIL responses, both by distinct regulatory features, and further imply that the differentiation state of malignant cells determines their sensitivity to death receptor signals.
Figures
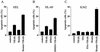

Similar articles
-
Human astrocytes are resistant to Fas ligand and tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis.J Neurosci. 2006 Mar 22;26(12):3299-308. doi: 10.1523/JNEUROSCI.5572-05.2006. J Neurosci. 2006. PMID: 16554480 Free PMC article.
-
Cotreatment with histone deacetylase inhibitor LAQ824 enhances Apo-2L/tumor necrosis factor-related apoptosis inducing ligand-induced death inducing signaling complex activity and apoptosis of human acute leukemia cells.Cancer Res. 2004 Apr 1;64(7):2580-9. doi: 10.1158/0008-5472.can-03-2629. Cancer Res. 2004. PMID: 15059915
-
Fas-associated death domain protein (FADD) and caspase-8 mediate up-regulation of c-Fos by Fas ligand and tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) via a FLICE inhibitory protein (FLIP)-regulated pathway.J Biol Chem. 2001 Aug 31;276(35):32585-90. doi: 10.1074/jbc.M100444200. Epub 2001 May 30. J Biol Chem. 2001. PMID: 11384965
-
Mechanisms of resistance to TRAIL-induced apoptosis in cancer.Cancer Gene Ther. 2005 Mar;12(3):228-37. doi: 10.1038/sj.cgt.7700792. Cancer Gene Ther. 2005. PMID: 15550937 Review.
-
Regulation of TRAIL-induced apoptosis by ectopic expression of antiapoptotic factors.Vitam Horm. 2004;67:453-83. doi: 10.1016/S0083-6729(04)67023-3. Vitam Horm. 2004. PMID: 15110190 Review.
Cited by
-
Telomere-homologous G-rich oligonucleotides sensitize human ovarian cancer cells to TRAIL-induced growth inhibition and apoptosis.Nucleic Acid Ther. 2013 Jun;23(3):167-74. doi: 10.1089/nat.2012.0401. Epub 2013 May 2. Nucleic Acid Ther. 2013. PMID: 23634944 Free PMC article.
-
Granzyme H destroys the function of critical adenoviral proteins required for viral DNA replication and granzyme B inhibition.EMBO J. 2007 Apr 18;26(8):2148-57. doi: 10.1038/sj.emboj.7601650. Epub 2007 Mar 15. EMBO J. 2007. PMID: 17363894 Free PMC article.
-
Inability of plasmacytoid dendritic cells to directly lyse HIV-infected autologous CD4+ T cells despite induction of tumor necrosis factor-related apoptosis-inducing ligand.J Virol. 2010 Mar;84(6):2762-73. doi: 10.1128/JVI.01350-09. Epub 2009 Dec 30. J Virol. 2010. PMID: 20042498 Free PMC article.
-
The scavenging of superoxide radicals promotes apoptosis induced by a novel cell-permeable fusion protein, sTRAIL:FeSOD, in tumor necrosis factor-related apoptosis-inducing ligand-resistant leukemia cells.BMC Biol. 2011 Mar 19;9:18. doi: 10.1186/1741-7007-9-18. BMC Biol. 2011. PMID: 21418589 Free PMC article.
-
Activation of the Akt survival pathway contributes to TRAIL resistance in cancer cells.PLoS One. 2010 Apr 19;5(4):e10226. doi: 10.1371/journal.pone.0010226. PLoS One. 2010. PMID: 20419107 Free PMC article.
References
-
- Altucci, L., A. Rossin, W. Raffelsberger, A. Reitmair, C. Chomienne, and H. Gronemeyer. 2001. Retinoic acid-induced apoptosis in leukemia cells is mediated by paracrine action of tumor-selective death ligand TRAIL. Nat. Med. 7:680-686. - PubMed
-
- Andersson, L. C., M. Jokinen, and C. G. Gahmberg. 1979. Induction of erythroid differentiation in the human leukaemia cell line K562. Nature 278:364-365. - PubMed
-
- Ashkenazi, A., R. C. Pai, S. Fong, S. Leung, D. A. Lawrence, S. A. Marsters, C. Blackie, L. Chang, A. E. McMurtrey, A. Hebert, L. DeForge, I. L. Koumenis, D. Lewis, L. Harris, J. Bussiere, H. Koeppen, Z. Shahrokh, and R. H. Schwall. 1999. Safety and antitumor activity of recombinant soluble Apo2 ligand. J. Clin. Investig. 104:155-162. - PMC - PubMed
-
- Benito, A., M. Silva, D. Grillot, G. Nunez, and J. L. Fernandez-Luna. 1996. Apoptosis induced by erythroid differentiation of human leukemia cell lines is inhibited by Bcl-XL. Blood 87:3837-3843. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
