Kep1 interacts genetically with dredd/caspase-8, and kep1 mutants alter the balance of dredd isoforms
- PMID: 12563030
- PMCID: PMC149916
- DOI: 10.1073/pnas.0236048100
Kep1 interacts genetically with dredd/caspase-8, and kep1 mutants alter the balance of dredd isoforms
Abstract
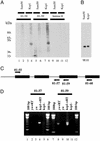
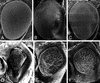
The Drosophila kep1 gene encodes an RNA binding protein related to the murine QUAKING apoptotic inducer. We have previously shown that kep1 can induce apoptosis when transfected into different cell lines. To better define the role of Kep1 in apoptosis, we generated kep1 null flies. These flies were viable, but females displayed reduced fertility, with approximately half of the eggs laid from kep1- homozygotes failing to hatch. In addition, loss of kep1 suppressed GMR-rpr-mediated apoptosis in the Drosophila eye, and kep1 mutant flies had increased susceptibility to Escherichia coli infection. We found that Kep1 bound dredd RNA in vitro, and that extracts prepared from kep1 mutant ovaries had markedly reduced proteolytic cleavage activity toward the caspase-8 target substrate IETD-7-amino-4-trifluoromethyl coumarin. We observed increased levels of the beta isoform of dredd mRNA in kep1 mutants as compared with wild-type. Taken together, our results suggest that Kep1 regulates apoptosis by influencing the processing of dredd RNA.
Figures



Similar articles
-
Phosphorylation status of the Kep1 protein alters its affinity for its protein binding partner alternative splicing factor ASF/SF2.Biochem J. 2006 Nov 15;400(1):91-7. doi: 10.1042/BJ20060384. Biochem J. 2006. PMID: 16834570 Free PMC article.
-
Targeted mutagenesis of the Sap47 gene of Drosophila: flies lacking the synapse associated protein of 47 kDa are viable and fertile.BMC Neurosci. 2004 Apr 29;5:16. doi: 10.1186/1471-2202-5-16. BMC Neurosci. 2004. PMID: 15117418 Free PMC article.
-
dXNP, a Drosophila homolog of XNP/ATRX, induces apoptosis via Jun-N-terminal kinase activation.FEBS Lett. 2007 Jun 12;581(14):2625-32. doi: 10.1016/j.febslet.2007.05.005. Epub 2007 May 11. FEBS Lett. 2007. PMID: 17531976
-
Morphological irregularities and features of resistance to apoptosis in the dcp-1/pita double mutated egg chambers during Drosophila oogenesis.Cell Motil Cytoskeleton. 2005 Jan;60(1):14-23. doi: 10.1002/cm.20043. Cell Motil Cytoskeleton. 2005. PMID: 15547953
-
Drosophila star proteins: what can be learned from flies?Adv Exp Med Biol. 2010;693:93-105. Adv Exp Med Biol. 2010. PMID: 21189688 Review.
Cited by
-
Polydnavirus Ank proteins bind NF-κB homodimers and inhibit processing of Relish.PLoS Pathog. 2012;8(5):e1002722. doi: 10.1371/journal.ppat.1002722. Epub 2012 May 24. PLoS Pathog. 2012. PMID: 22654665 Free PMC article.
-
Specificity of the STAR/GSG domain protein Qk1: implications for the regulation of myelination.RNA. 2004 Sep;10(9):1449-58. doi: 10.1261/rna.7780504. Epub 2004 Jul 23. RNA. 2004. PMID: 15273320 Free PMC article.
-
Phosphorylation status of the Kep1 protein alters its affinity for its protein binding partner alternative splicing factor ASF/SF2.Biochem J. 2006 Nov 15;400(1):91-7. doi: 10.1042/BJ20060384. Biochem J. 2006. PMID: 16834570 Free PMC article.
-
Evolutionary Dynamics of GLD-1-mRNA complexes in Caenorhabditis nematodes.Genome Biol Evol. 2014 Dec 9;7(1):314-35. doi: 10.1093/gbe/evu272. Genome Biol Evol. 2014. PMID: 25502909 Free PMC article.
-
A young Drosophila duplicate gene plays essential roles in spermatogenesis by regulating several Y-linked male fertility genes.PLoS Genet. 2010 Dec 23;6(12):e1001255. doi: 10.1371/journal.pgen.1001255. PLoS Genet. 2010. PMID: 21203494 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

