The budding yeast Ipl1/Aurora protein kinase regulates mitotic spindle disassembly
- PMID: 12566427
- PMCID: PMC2172676
- DOI: 10.1083/jcb.200209018
The budding yeast Ipl1/Aurora protein kinase regulates mitotic spindle disassembly
Abstract
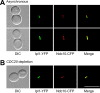
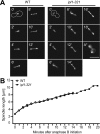
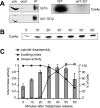
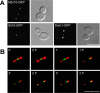
Ipl1p is the budding yeast member of the Aurora family of protein kinases, critical regulators of genomic stability that are required for chromosome segregation, the spindle checkpoint, and cytokinesis. Using time-lapse microscopy, we found that Ipl1p also has a function in mitotic spindle disassembly that is separable from its previously identified roles. Ipl1-GFP localizes to kinetochores from G1 to metaphase, transfers to the spindle after metaphase, and accumulates at the spindle midzone late in anaphase. Ipl1p kinase activity increases at anaphase, and ipl1 mutants can stabilize fragile spindles. As the spindle disassembles, Ipl1p follows the plus ends of the depolymerizing spindle microtubules. Many Ipl1p substrates colocalize with Ipl1p to the spindle midzone, identifying additional proteins that may regulate spindle disassembly. We propose that Ipl1p regulates both the kinetochore and interpolar microtubule plus ends to regulate its various mitotic functions.
Figures


References
-
- Adams, R.R., S.P. Wheatleya, A.M. Gouldsworthy, S.E. Kandels-Lewis, M. Carmena, C. Smythe, D.L. Gerloff, and W.C. Earnshaw. 2000. INCENP binds the aurora-related kinase AIRK2 and is required to target it to chromosomes, the central spindle and cleavage furrow. Curr. Biol. 10:1075–1078. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

