Lysine-79 of histone H3 is hypomethylated at silenced loci in yeast and mammalian cells: a potential mechanism for position-effect variegation
- PMID: 12574507
- PMCID: PMC149917
- DOI: 10.1073/pnas.0437846100
Lysine-79 of histone H3 is hypomethylated at silenced loci in yeast and mammalian cells: a potential mechanism for position-effect variegation
Abstract
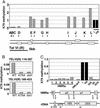
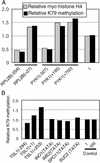
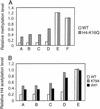
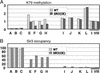
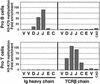
Methylation of lysine-79 (K79) within the globular domain of histone H3 by Dot1 methylase is important for transcriptional silencing and for association of the Sir silencing proteins in yeast. Here, we show that the level of H3-K79 methylation is low at all Sir-dependent silenced loci but not at other transcriptionally repressed regions. Hypomethylation of H3-K79 at the telomeric and silent mating-type loci, but not the ribosomal DNA, requires the Sir proteins. Overexpression of Sir3 concomitantly extends the domain of Sir protein association and H3-K79 hypomethylation at telomeres. In mammalian cells, H3-K79 methylation is found at loci that are active for V(D)J recombination, but not at recombinationally inactive loci that are heterochromatic. These results suggest that H3-K79 methylation is an evolutionarily conserved marker of active chromatin regions, and that silencing proteins block the ability of Dot1 to methylate histone H3. Further, they suggest that Sir proteins preferentially bind chromatin with hypomethylated H3-K79 and then block H3-K79 methylation. This positive feedback loop, and the reverse loop in which H3-K79 methylation weakens Sir protein association and leads to further methylation, suggests a model for position-effect variegation.
Figures

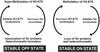
Similar articles
-
Insights into the impact of histone acetylation and methylation on Sir protein recruitment, spreading, and silencing in Saccharomyces cerevisiae.J Mol Biol. 2008 Sep 12;381(4):826-44. doi: 10.1016/j.jmb.2008.06.059. Epub 2008 Jun 28. J Mol Biol. 2008. PMID: 18619469
-
Symmetry, asymmetry, and kinetics of silencing establishment in Saccharomyces cerevisiae revealed by single-cell optical assays.Proc Natl Acad Sci U S A. 2011 Jan 25;108(4):1209-16. doi: 10.1073/pnas.1018742108. Epub 2011 Jan 24. Proc Natl Acad Sci U S A. 2011. PMID: 21262833 Free PMC article.
-
Lysine methylation within the globular domain of histone H3 by Dot1 is important for telomeric silencing and Sir protein association.Genes Dev. 2002 Jun 15;16(12):1518-27. doi: 10.1101/gad.1001502. Genes Dev. 2002. PMID: 12080090 Free PMC article.
-
[The roles of histone lysine methylation in epigenetic regulation].Yi Chuan. 2007 Apr;29(4):387-92. doi: 10.1360/yc-007-0387. Yi Chuan. 2007. PMID: 17548299 Review. Chinese.
-
Cross-talking histones: implications for the regulation of gene expression and DNA repair.Biochem Cell Biol. 2005 Aug;83(4):460-7. doi: 10.1139/o05-116. Biochem Cell Biol. 2005. PMID: 16094449 Review.
Cited by
-
The Ino80 complex prevents invasion of euchromatin into silent chromatin.Genes Dev. 2015 Feb 15;29(4):350-5. doi: 10.1101/gad.256255.114. Genes Dev. 2015. PMID: 25691465 Free PMC article.
-
Barrier proteins remodel and modify chromatin to restrict silenced domains.Mol Cell Biol. 2004 Mar;24(5):1956-67. doi: 10.1128/MCB.24.5.1956-1967.2004. Mol Cell Biol. 2004. PMID: 14966276 Free PMC article.
-
The ubiquitin-conjugating enzyme HR6B is required for maintenance of X chromosome silencing in mouse spermatocytes and spermatids.BMC Genomics. 2010 Jun 10;11:367. doi: 10.1186/1471-2164-11-367. BMC Genomics. 2010. PMID: 20537150 Free PMC article.
-
Assembly and analysis of the mouse immunoglobulin kappa gene sequence.Immunogenetics. 2004 Oct;56(7):490-505. doi: 10.1007/s00251-004-0659-0. Epub 2004 Sep 18. Immunogenetics. 2004. PMID: 15378297
-
Histone variant H2A.Z marks the 5' ends of both active and inactive genes in euchromatin.Cell. 2005 Oct 21;123(2):233-48. doi: 10.1016/j.cell.2005.10.002. Cell. 2005. PMID: 16239142 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

