Exonuclease-deficient polymerase mutant of herpes simplex virus type 1 induces altered spectra of mutations
- PMID: 12584319
- PMCID: PMC149784
- DOI: 10.1128/jvi.77.5.2946-2955.2003
Exonuclease-deficient polymerase mutant of herpes simplex virus type 1 induces altered spectra of mutations
Abstract
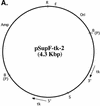
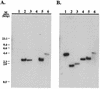
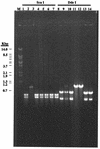
The effect of exonuclease activity of the herpes simplex virus DNA polymerase (Pol) on DNA replication fidelity was examined by using the supF mutagenesis assay. The recombinants with exonuclease-deficient Pol, containing an integrated supF gene in the thymidine kinase locus (tk), exhibited supF mutation frequencies ranging from 0.14 to 5.6%, consistent with the tk mutation frequencies reported previously (Y. T. Hwang, B.-Y. Liu, D. M. Coen, and C. B. C. Hwang, J. Virol. 71:7791-7798, 1997). The increased mutation frequencies were 10- to 500-fold higher than those observed for wild-type Pol recombinants. The increased mutation frequencies also were significantly higher than those of supF mutant replicated by exonuclease-deficient Pols in the plasmid-borne assay. Furthermore, characterization of supF mutants demonstrated that recombinants with a defective exonuclease induced types and distributions of supF mutations different from those induced by wild-type Pol recombinants. The types of supF mutations induced by exonuclease-deficient recombinants differed between the plasmid- and genome-based assays. The spectra of supF mutations also differed between the two assays. In addition, exonuclease-defective viruses also induced different spectra of supF and tk mutations. Therefore, both the assay methods and the target genes used for mutagenesis studies can affect the repication fidelity of herpes simplex virus type 1 Pol with defective exonuclease activity.
Figures


Similar articles
-
Replication fidelity of the supF gene integrated in the thymidine kinase locus of herpes simplex virus type 1.J Virol. 2002 Apr;76(8):3605-14. doi: 10.1128/jvi.76.8.3605-3614.2002. J Virol. 2002. PMID: 11907200 Free PMC article.
-
Effects of exonuclease activity and nucleotide selectivity of the herpes simplex virus DNA polymerase on the fidelity of DNA replication in vivo.J Virol. 1999 Jul;73(7):5326-32. doi: 10.1128/JVI.73.7.5326-5332.1999. J Virol. 1999. PMID: 10364279 Free PMC article.
-
Assessing the contribution of the herpes simplex virus DNA polymerase to spontaneous mutations.BMC Infect Dis. 2002 May 7;2:7. doi: 10.1186/1471-2334-2-7. BMC Infect Dis. 2002. PMID: 12019036 Free PMC article.
-
Mutation spectra in supF: approaches to elucidating sequence context effects.Mutat Res. 2000 May 30;450(1-2):61-73. doi: 10.1016/s0027-5107(00)00016-6. Mutat Res. 2000. PMID: 10838134 Review.
-
Mutations induced by reactive nitrogen oxide species in the supF forward mutation assay.Mutat Res. 2000 May 30;450(1-2):95-105. doi: 10.1016/s0027-5107(00)00018-x. Mutat Res. 2000. PMID: 10838136 Review.
Cited by
-
Clusters of mutations from transient hypermutability.Proc Natl Acad Sci U S A. 2005 Sep 6;102(36):12849-54. doi: 10.1073/pnas.0503009102. Epub 2005 Aug 23. Proc Natl Acad Sci U S A. 2005. PMID: 16118275 Free PMC article.
-
Two-Metal Ion-Dependent Enzymes as Potential Antiviral Targets in Human Herpesviruses.mBio. 2022 Feb 22;13(1):e0322621. doi: 10.1128/mbio.03226-21. Epub 2022 Jan 25. mBio. 2022. PMID: 35073739 Free PMC article.
-
Too many mutants with multiple mutations.Crit Rev Biochem Mol Biol. 2007 Jul-Aug;42(4):247-58. doi: 10.1080/10409230701495631. Crit Rev Biochem Mol Biol. 2007. PMID: 17687667 Free PMC article. Review.
-
Viral Nucleases from Herpesviruses and Coronavirus in Recombination and Proofreading: Potential Targets for Antiviral Drug Discovery.Viruses. 2022 Jul 16;14(7):1557. doi: 10.3390/v14071557. Viruses. 2022. PMID: 35891537 Free PMC article. Review.
-
A point mutation within conserved region VI of herpes simplex virus type 1 DNA polymerase confers altered drug sensitivity and enhances replication fidelity.J Virol. 2004 Jan;78(2):650-7. doi: 10.1128/jvi.78.2.650-657.2004. J Virol. 2004. PMID: 14694096 Free PMC article.
References
-
- Chiou, H. C., K. Kumura, A. Hu, K. M. Kern, and D. M. Coen. 1995. Penciclovir-resistance mutations in the herpes simplex virus DNA polymerase gene. Antivir. Chem. Chemother. 6:281-288.
-
- Clarke, G. M. 1994. Statistics and experimental design. An introduction for biologists and biochemists, 3rd ed., p. 82-83. Edward Arnold, London, England.
-
- Coen, D. M. 1996. Viral DNA polymerases, p. 495-523. In M. L. DePamphilis (ed.), DNA replication in eukaryotic cells. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
-
- Fisher, L. D., and G. van Belle. 1993. Biostatistics—a methodology for the health sciences, p. 218-221. John Wiley & Sons, Inc., New York, N.Y.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

