Subcellular localization of herpes simplex virus type 1 UL51 protein and role of palmitoylation in Golgi apparatus targeting
- PMID: 12584344
- PMCID: PMC149782
- DOI: 10.1128/jvi.77.5.3204-3216.2003
Subcellular localization of herpes simplex virus type 1 UL51 protein and role of palmitoylation in Golgi apparatus targeting
Abstract
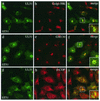
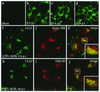
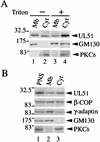
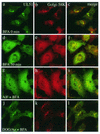
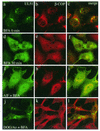
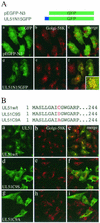
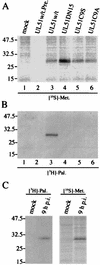
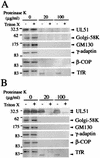
The herpes simplex virus type 1 (HSV-1) UL51 gene products are virion-associated phosphoproteins with apparent molecular masses of 27, 29, and 30 kDa in HSV-1-infected cells. In this study, we have investigated the intracellular localization and distribution of UL51 protein both in infected cells and in transfected cells expressing only UL51. We found that this protein colocalized closely with Golgi marker proteins such as the Golgi-58K protein and GM130 in transfected cells expressing only UL51. However, in infected cells, the UL51 protein localized to the juxtanuclear region but only partially colocalized with the Golgi maker proteins. Mutant protein analysis revealed that the N-terminal 15 amino acid residues of the UL51 protein sufficed for this Golgi localization property. The UL51 protein redistributed on addition of brefeldin A. This was prevented by pretreatment with 2-deoxyglucose and sodium azide, which results in ATP depletion, but not by pretreatment with NaF and AlCl(3), which activates heterotrimeric G proteins. Moreover, we found that palmitoylation of the UL51 protein through the N-terminal cysteine at position 9 was necessary for its Golgi localization. Protease digestion analysis suggested that the UL51 protein localized on the cytoplasmic face of the membrane in UL51-transfected cells, while in infected cells it localized mainly to the inside of cytoplasmic vesicles and/or the viral envelope. Transmission immunoelectron microscopy revealed an association of UL51 protein-specific labeling with cytoplasmic virions and also with some membranous structure. We infer from these observations that internalization of UL51 protein into the cytoplasmic vesicle and/or virion may occur in association with viral envelopment in HSV-infected cells.
Figures



Similar articles
-
Herpes simplex virus type 1 UL51 protein is involved in maturation and egress of virus particles.J Virol. 2005 Jun;79(11):6947-56. doi: 10.1128/JVI.79.11.6947-6956.2005. J Virol. 2005. PMID: 15890934 Free PMC article.
-
The Interaction between Herpes Simplex Virus 1 Tegument Proteins UL51 and UL14 and Its Role in Virion Morphogenesis.J Virol. 2016 Sep 12;90(19):8754-67. doi: 10.1128/JVI.01258-16. Print 2016 Oct 1. J Virol. 2016. PMID: 27440890 Free PMC article.
-
Characterization of subcellular localization of duck enteritis virus UL51 protein.Virol J. 2009 Jul 3;6:92. doi: 10.1186/1743-422X-6-92. Virol J. 2009. PMID: 19575796 Free PMC article.
-
Identification and characterization of the herpes simplex virus type 1 UL51 gene product.J Gen Virol. 1998 Dec;79 ( Pt 12):3027-31. doi: 10.1099/0022-1317-79-12-3027. J Gen Virol. 1998. PMID: 9880018
-
Functions of the UL51 protein during the herpesvirus life cycle.Front Microbiol. 2024 Aug 26;15:1457582. doi: 10.3389/fmicb.2024.1457582. eCollection 2024. Front Microbiol. 2024. PMID: 39252835 Free PMC article. Review.
Cited by
-
Palmitoylation of virus proteins.Biol Cell. 2012 Sep;104(9):493-515. doi: 10.1111/boc.201200006. Epub 2012 Jul 17. Biol Cell. 2012. PMID: 22548323 Free PMC article. Review.
-
Packaging determinants in the UL11 tegument protein of herpes simplex virus type 1.J Virol. 2006 Nov;80(21):10534-41. doi: 10.1128/JVI.01172-06. Epub 2006 Aug 23. J Virol. 2006. PMID: 16928743 Free PMC article.
-
Importance of conserved cysteine residues in the coronavirus envelope protein.J Virol. 2008 Mar;82(6):3000-10. doi: 10.1128/JVI.01914-07. Epub 2008 Jan 9. J Virol. 2008. PMID: 18184703 Free PMC article.
-
A Tyrosine-Based Trafficking Motif of the Tegument Protein pUL71 Is Crucial for Human Cytomegalovirus Secondary Envelopment.J Virol. 2017 Dec 14;92(1):e00907-17. doi: 10.1128/JVI.00907-17. Print 2018 Jan 1. J Virol. 2017. PMID: 29046458 Free PMC article.
-
Herpes simplex virus type 1 UL51 protein is involved in maturation and egress of virus particles.J Virol. 2005 Jun;79(11):6947-56. doi: 10.1128/JVI.79.11.6947-6956.2005. J Virol. 2005. PMID: 15890934 Free PMC article.
References
-
- Baer, R. J., A. T. Bankier, M. D. Biggin, P. L. Deininger, P. J. Farrell, T. J. Gibson, G. Hatfull, G. S. Hudson, S. C. Satchwell, C. Seguin, P. Tuffnell, and B. G. Barrell. 1984. DNA sequence and expression of the B95-8 Epstein-Barr virus genome. Nature (London) 310:207-211. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

