Structural origins of adenine-tract bending
- PMID: 12586860
- PMCID: PMC151347
- DOI: 10.1073/pnas.0437877100
Structural origins of adenine-tract bending
Abstract
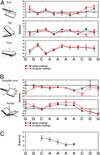
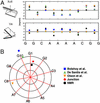
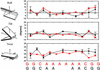
DNA sequences containing short adenine tracts are intrinsically curved and play a role in transcriptional regulation. Despite many high-resolution NMR and x-ray studies, the origins of curvature remain disputed. Long-range restraints provided by 85 residual dipolar couplings were measured for a DNA decamer containing an adenine (A)(4)-tract and used to refine the structure. The overall bend in the molecule is a result of in-phase negative roll in the A-tract and positive roll at its 5' junction, as well as positive and negative tilt inside the A-tract and near its junctions. The bend magnitude and direction obtained from NMR structures is 9.0 degrees into the minor groove in a coordinate frame located at the third AT base pair. We evaluated long-range and wedge models for DNA curvature and concluded that our data for A-tract curvature are best explained by a "delocalized bend" model. The global bend magnitude and direction of the NMR structure are in excellent agreement with the junction model parameters used to rationalize gel electrophoretic data and with preliminary results of a cyclization kinetics assay from our laboratory.
Figures

References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

