Chemical genetic analysis of Apg1 reveals a non-kinase role in the induction of autophagy
- PMID: 12589048
- PMCID: PMC149986
- DOI: 10.1091/mbc.e02-07-0413
Chemical genetic analysis of Apg1 reveals a non-kinase role in the induction of autophagy
Abstract
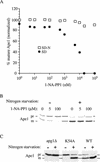
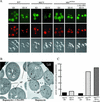
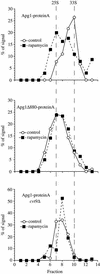
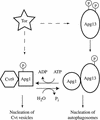
Macroautophagy is a catabolic membrane trafficking phenomenon that is observed in all eukaryotic cells in response to various stimuli, such as nitrogen starvation and challenge with specific hormones. In the yeast Saccharomyces cerevisiae, the induction of autophagy involves a direct signal transduction mechanism that affects membrane dynamics. In this system, the induction process modifies a constitutive trafficking pathway called the cytoplasm-to-vacuole targeting (Cvt) pathway, which transports the vacuolar hydrolase aminopeptidase I, from the formation of small Cvt vesicles to the formation of autophagosomes. Apg1 is one of the proteins required for the direct signal transduction cascade that modifies membrane dynamics. Although Apg1 is required for both the Cvt pathway and autophagy, we find that Apg1 kinase activity is required only for Cvt trafficking of aminopeptidase I but not for import via autophagy. In addition, the data support a novel role for Apg1 in nucleation of autophagosomes that is distinct from its catalytic kinase activity and imply a qualitative difference in the mechanism of autophagosome and Cvt vesicle formation.
Figures




References
-
- Abeliovich H, Grote E, Novick P, Ferro-Novick S. Tlg2p, a yeast syntaxin homolog that resides on the Golgi and Endocytic structures. J Biol Chem. 1998;273:11719–11727. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

