Discovery and characterization of Acanthamoeba castellanii mitochondrial 5S rRNA
- PMID: 12592002
- PMCID: PMC1370395
- DOI: 10.1261/rna.2170803
Discovery and characterization of Acanthamoeba castellanii mitochondrial 5S rRNA
Abstract
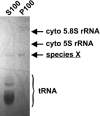
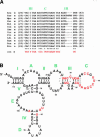
Although 5S rRNA is a highly conserved and universal component of eubacterial, archaeal, chloroplast, and eukaryotic cytoplasmic ribosomes, a mitochondrial DNA-encoded 5S rRNA has so far been identified only in land plants and certain protists. This raises the question of whether 5S rRNA is actually required for and used in mitochondrial translation. In the protist Acanthamoeba castellanii, BLAST searches fail to reveal a 5S rRNA gene in the complete mitochondrial genome sequence, nor is a 5S-sized RNA species detectable in ethidium bromide-stained gels of highly purified mitochondrial RNA preparations. Here we show that an alternative visualization technique, UV shadowing, readily detects a novel, mitochondrion-specific small RNA in A. castellanii mitochondrial RNA preparations, and that this RNA species is, in fact, a 5S rRNA encoded by the A. castellanii mitochondrial genome. These results emphasize the need for caution when interpreting negative results that suggest the absence of 5S rRNA and/or a mitochondrial DNA-encoded 5S rRNA sequence in other (particularly protist) mitochondrial systems.
Figures


Similar articles
-
Abundant 5S rRNA-like transcripts encoded by the mitochondrial genome in amoebozoa.Eukaryot Cell. 2010 May;9(5):762-73. doi: 10.1128/EC.00013-10. Epub 2010 Mar 19. Eukaryot Cell. 2010. PMID: 20304999 Free PMC article.
-
The ribosomal RNA gene region in Acanthamoeba castellanii mitochondrial DNA. A case of evolutionary transfer of introns between mitochondria and plastids?J Mol Biol. 1994 Jun 17;239(4):476-99. doi: 10.1006/jmbi.1994.1390. J Mol Biol. 1994. PMID: 8006963
-
Sequence and organization of 5S RNA genes from the eukaryotic protist Acanthamoeba castellanii.Gene. 1991 May 15;101(1):153-7. doi: 10.1016/0378-1119(91)90239-8. Gene. 1991. PMID: 1676384
-
Genes of Acanthamoeba: DNA, RNA and protein sequences (a review).J Protozool. 1990 Jul-Aug;37(4):17S-25S. doi: 10.1111/j.1550-7408.1990.tb01141.x. J Protozool. 1990. PMID: 1701831 Review.
-
5S rRNA and ribosome.Biochemistry (Mosc). 2011 Dec;76(13):1450-64. doi: 10.1134/S0006297911130062. Biochemistry (Mosc). 2011. PMID: 22339598 Review.
Cited by
-
Fragmentation of the large subunit ribosomal RNA gene in oyster mitochondrial genomes.BMC Genomics. 2010 Sep 2;11:485. doi: 10.1186/1471-2164-11-485. BMC Genomics. 2010. PMID: 20813041 Free PMC article.
-
Extreme RNA editing in coding islands and abundant microsatellites in repeat sequences of Selaginella moellendorffii mitochondria: the root of frequent plant mtDNA recombination in early tracheophytes.Genome Biol Evol. 2011;3:344-58. doi: 10.1093/gbe/evr027. Epub 2011 Mar 23. Genome Biol Evol. 2011. PMID: 21436122 Free PMC article.
-
Abundant 5S rRNA-like transcripts encoded by the mitochondrial genome in amoebozoa.Eukaryot Cell. 2010 May;9(5):762-73. doi: 10.1128/EC.00013-10. Epub 2010 Mar 19. Eukaryot Cell. 2010. PMID: 20304999 Free PMC article.
-
How to build a ribosome from RNA fragments in Chlamydomonas mitochondria.Nat Commun. 2021 Dec 9;12(1):7176. doi: 10.1038/s41467-021-27200-z. Nat Commun. 2021. PMID: 34887394 Free PMC article.
-
Characteristics of the nuclear (18S, 5.8S, 28S and 5S) and mitochondrial (12S and 16S) rRNA genes of Apis mellifera (Insecta: Hymenoptera): structure, organization, and retrotransposable elements.Insect Mol Biol. 2006 Oct;15(5):657-86. doi: 10.1111/j.1365-2583.2006.00689.x. Insect Mol Biol. 2006. PMID: 17069639 Free PMC article.
References
-
- Burger, G., Plante, I., Lonergan, K.M., and Gray, M.W. 1995. The mitochondrial DNA of the amoeboid protozoon, Acanthamoeba castellanii: Complete sequence, genome content and genome organization. J. Mol. Biol. 245: 522–537. - PubMed
-
- Cavalier-Smith, T. 1998. A revised six-kingdom system of life. Biol. Rev. Camb. Philos. Soc. 73: 203–266. - PubMed
-
- Cunningham, R.S., Bonen, L., Doolittle, W.F., and Gray, M.W. 1976. Unique species of 5 S 18 S and 26 S ribosomal RNA in wheat mitochondria. FEBS Lett. 69: 116–122. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
