Imprinting regulation of the murine Meg1/Grb10 and human GRB10 genes; roles of brain-specific promoters and mouse-specific CTCF-binding sites
- PMID: 12595547
- PMCID: PMC149825
- DOI: 10.1093/nar/gkg232
Imprinting regulation of the murine Meg1/Grb10 and human GRB10 genes; roles of brain-specific promoters and mouse-specific CTCF-binding sites
Abstract
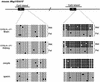
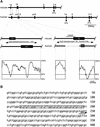
The imprinted mouse gene Meg1/Grb10 is expres sed from maternal alleles in almost all tissues and organs, except in the brain, where it is expressed biallelically, and the paternal allele is expressed preferentially in adulthood. In contrast, the human GRB10 gene shows equal biallelic expression in almost all tissues and organs, while it is almost always expressed paternally in the fetal brain. To elucidate the molecular mechanisms of the complex imprinting patterns among the different tissues and organs of humans and mice, we analyzed in detail both the genomic structures and tissue-specific expression profiles of these species. Experiments using 5'-RACE and RT-PCR demonstrated the existence in both humans and mice of novel brain- specific promoters, in which only the paternal allele was active. The promoters were located in the primary differentially methylated regions. Interest ingly, CTCF-binding sites were found only in the mouse promoter region where CTCF showed DNA methylation-sensitive binding activity. Thus, the insulator function of CTCF might cause reciprocal maternal expression of the Meg1/Grb10 gene from another upstream promoter in the mouse, whereas the human upstream promoter is active in both parental alleles due to the lack of the corresponding insulator sequence in this region.
Figures

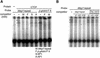

Similar articles
-
Role of DNA methylation and histone H3 lysine 27 methylation in tissue-specific imprinting of mouse Grb10.Mol Cell Biol. 2007 Jan;27(2):732-42. doi: 10.1128/MCB.01329-06. Epub 2006 Nov 13. Mol Cell Biol. 2007. PMID: 17101788 Free PMC article.
-
Conserved methylation imprints in the human and mouse GRB10 genes with divergent allelic expression suggests differential reading of the same mark.Hum Mol Genet. 2003 May 1;12(9):1005-19. doi: 10.1093/hmg/ddg110. Hum Mol Genet. 2003. PMID: 12700169
-
Paternal deletion of Meg1/Grb10 DMR causes maternalization of the Meg1/Grb10 cluster in mouse proximal Chromosome 11 leading to severe pre- and postnatal growth retardation.Hum Mol Genet. 2009 Apr 15;18(8):1424-38. doi: 10.1093/hmg/ddp049. Epub 2009 Jan 27. Hum Mol Genet. 2009. PMID: 19174477
-
Differential 3D chromatin organization and gene activity in genomic imprinting.Curr Opin Genet Dev. 2020 Apr;61:17-24. doi: 10.1016/j.gde.2020.03.004. Epub 2020 Apr 13. Curr Opin Genet Dev. 2020. PMID: 32299027 Review.
-
Tissue-specific regulation and function of Grb10 during growth and neuronal commitment.Proc Natl Acad Sci U S A. 2015 Jun 2;112(22):6841-7. doi: 10.1073/pnas.1411254111. Epub 2014 Nov 3. Proc Natl Acad Sci U S A. 2015. PMID: 25368187 Free PMC article. Review.
Cited by
-
Imprinted methylation profiles for proximal mouse chromosomes 11 and 7 as revealed by methylation-sensitive representational difference analysis.Mamm Genome. 2003 Dec;14(12):805-16. doi: 10.1007/s00335-003-2287-7. Mamm Genome. 2003. PMID: 14724735
-
Dnmt3a1 upregulates transcription of distinct genes and targets chromosomal gene clusters for epigenetic silencing in mouse embryonic stem cells.Mol Cell Biol. 2011 Apr;31(7):1577-92. doi: 10.1128/MCB.01093-10. Epub 2011 Jan 24. Mol Cell Biol. 2011. PMID: 21262766 Free PMC article.
-
Characterization of the imprinting signature of mouse embryo fibroblasts by RNA deep sequencing.Nucleic Acids Res. 2014 Feb;42(3):1772-83. doi: 10.1093/nar/gkt1042. Epub 2013 Nov 11. Nucleic Acids Res. 2014. PMID: 24217910 Free PMC article.
-
Susceptibility to astrocytoma in mice mutant for Nf1 and Trp53 is linked to chromosome 11 and subject to epigenetic effects.Proc Natl Acad Sci U S A. 2004 Aug 31;101(35):13008-13. doi: 10.1073/pnas.0401236101. Epub 2004 Aug 19. Proc Natl Acad Sci U S A. 2004. PMID: 15319471 Free PMC article.
-
Real-time PCR analysis of candidate imprinted genes on mouse chromosome 11 shows balanced expression from the maternal and paternal chromosomes and strain-specific variation in expression levels.Epigenetics. 2008 Jan-Feb;3(1):43-50. doi: 10.4161/epi.3.1.5469. Epub 2007 Dec 20. Epigenetics. 2008. PMID: 18188004 Free PMC article.
References
-
- Miyoshi N., Kuroiwa,Y., Kohda,T., Shitara,H., Yonekawa,H., Kawabe,T., Hasegawa,H., Barton,S.C., Surani,M.A., Kaneko-Ishino,T. and Ishino,F. (1998) Identification of the Meg1/Grb10 imprinted gene on mouse proximal chromosome 11, a candidate for the Silver-Russell syndrome gene. Proc. Natl Acad. Sci. USA, 95, 1102–1107. - PMC - PubMed
-
- Bai R.Y., Jahn,T., Schrem,S., Munzert,G., Weidner,K.M., Wang,J.Y. and Duyster,J. (1998) The SH2-containing adapter protein GRB10 interacts with BCR-ABL. Oncogene, 17, 941–948. - PubMed
-
- O’Neill T.J., Rose,D.W., Pillay,T.S., Hotta,K., Olefsky,J.M. and Gustafson,T.A. (1996)) Interaction of a GRB-IR splice variant (a human GRB10 homolog) with the insulin and insulin-like growth factor I receptors. Evidence for a role in mitogenic signaling. J. Biol. Chem., 271, 22506–22513. - PubMed
-
- Morrione A., Valentinis,B., Resnicoff,M., Xu,S. and Baserga,R. (1997) The role of mGrb10alpha in insulin-like growth factor I-mediated growth. J. Biol. Chem., 272, 26382–26387. - PubMed
-
- Liu F. and Roth,R.A. (1998) Binding of SH2 containing proteins to the insulin receptor: a new way for modulating insulin signalling. Mol. Cell. Biochem., 182, 73–78. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

