Molecular typing of IberoAmerican Cryptococcus neoformans isolates
- PMID: 12603989
- PMCID: PMC2901947
- DOI: 10.3201/eid0902.020246
Molecular typing of IberoAmerican Cryptococcus neoformans isolates
Abstract
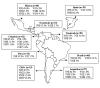
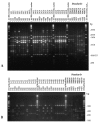
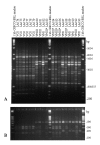
A network was established to acquire basic knowledge of Cryptococcus neoformans in IberoAmerican countries. To this effect, 340 clinical, veterinary, and environmental isolates from Argentina, Brazil, Chile, Colombia, Mexico, Peru, Venezuela, Guatemala, and Spain were typed by using M13 polymerase chain reaction-fingerprinting and orotidine monophosphate pyrophosphorylase (URA5) gene restriction fragment length polymorphism analysis with HhaI and Sau96I in a double digest. Both techniques grouped all isolates into eight previously established molecular types. The majority of the isolates, 68.2% (n=232), were VNI (var. grubii, serotype A), which accords with the fact that this variety causes most human cryptococcal infections worldwide. A smaller proportion, 5.6% (n=19), were VNII (var. grubii, serotype A); 4.1% (n=14), VNIII (AD hybrid), with 9 isolates having a polymorphism in the URA5 gene; 1.8% (n=6), VNIV (var. neoformans, serotype D); 3.5% (n=12), VGI; 6.2% (n=21), VGII; 9.1% (n=31), VGIII, and 1.5% (n=5) VGIV, with all four VG types containing var. gatii serotypes B and C isolates.
Figures

References
-
- Casadevall A, Perfect JR. Cryptococcus neoformans. Washington: ASM Press; 1998.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
