Localization of DNA polymerases eta and iota to the replication machinery is tightly co-ordinated in human cells
- PMID: 12606586
- PMCID: PMC150329
- DOI: 10.1093/emboj/cdf618
Localization of DNA polymerases eta and iota to the replication machinery is tightly co-ordinated in human cells
Abstract
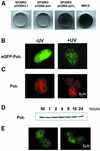
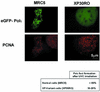
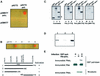
Y-family DNA polymerases can replicate past a variety of damaged bases in vitro but, with the exception of DNA polymerase eta (poleta), which is defective in xeroderma pigmentosum variants, there is little information on the functions of these polymerases in vivo. Here, we show that DNA polymerase iota (poliota), like poleta, associates with the replication machinery and accumulates at stalled replication forks following DNA-damaging treatment. We show that poleta and poliota foci form with identical kinetics and spatial distributions, suggesting that localization of these two polymerases is tightly co-ordinated within the nucleus. Furthermore, localization of poliota in replication foci is largely dependent on the presence of poleta. Using several different approaches, we demonstrate that poleta and poliota interact with each other physically and that the C-terminal 224 amino acids of poliota are sufficient for both the interaction with poleta and accumulation in replication foci. Our results provide strong evidence that poleta targets poliota to the replication machinery, where it may play a general role in maintaining genome integrity as well as participating in translesion DNA synthesis.
Figures



Corrected and republished from
-
Localization of DNA polymerases eta and iota to the replication machinery is tightly co-ordinated in human cells.EMBO J. 2002 Nov 15;21(22):6246-56. doi: 10.1093/emboj/cdf618. EMBO J. 2002. Corrected and republished in: EMBO J. 2003 Mar 3;22(5):1223-33. doi: 10.1093/emboj/cdf618. PMID: 12426396 Free PMC article. Corrected and republished.
Similar articles
-
Localization of DNA polymerases eta and iota to the replication machinery is tightly co-ordinated in human cells.EMBO J. 2002 Nov 15;21(22):6246-56. doi: 10.1093/emboj/cdf618. EMBO J. 2002. Corrected and republished in: EMBO J. 2003 Mar 3;22(5):1223-33. doi: 10.1093/emboj/cdf618. PMID: 12426396 Free PMC article. Corrected and republished.
-
Proliferating cell nuclear antigen-dependent coordination of the biological functions of human DNA polymerase iota.J Biol Chem. 2004 Nov 12;279(46):48360-8. doi: 10.1074/jbc.M406511200. Epub 2004 Sep 1. J Biol Chem. 2004. PMID: 15342632
-
Effect of proliferating cell nuclear antigen ubiquitination and chromatin structure on the dynamic properties of the Y-family DNA polymerases.Mol Biol Cell. 2008 Dec;19(12):5193-202. doi: 10.1091/mbc.e08-07-0724. Epub 2008 Sep 17. Mol Biol Cell. 2008. PMID: 18799611 Free PMC article.
-
poliota-dependent lesion bypass in vitro.Mutat Res. 2002 Dec 29;510(1-2):9-22. doi: 10.1016/s0027-5107(02)00248-8. Mutat Res. 2002. PMID: 12459439 Review.
-
Replication of damaged DNA in mammalian cells: new solutions to an old problem.Mutat Res. 2002 Nov 30;509(1-2):23-34. doi: 10.1016/s0027-5107(02)00227-0. Mutat Res. 2002. PMID: 12427529 Review.
Cited by
-
Division of labor of Y-family polymerases in translesion-DNA synthesis for distinct types of DNA damage.PLoS One. 2021 Jun 1;16(6):e0252587. doi: 10.1371/journal.pone.0252587. eCollection 2021. PLoS One. 2021. PMID: 34061890 Free PMC article.
-
Post-Translational Modifications of PCNA: Guiding for the Best DNA Damage Tolerance Choice.J Fungi (Basel). 2022 Jun 10;8(6):621. doi: 10.3390/jof8060621. J Fungi (Basel). 2022. PMID: 35736104 Free PMC article. Review.
-
Translesion DNA synthesis in the context of cancer research.Cancer Cell Int. 2011 Nov 2;11:39. doi: 10.1186/1475-2867-11-39. Cancer Cell Int. 2011. PMID: 22047021 Free PMC article.
-
The Response of the Replication Apparatus to Leading Template Strand Blocks.Cells. 2023 Nov 11;12(22):2607. doi: 10.3390/cells12222607. Cells. 2023. PMID: 37998342 Free PMC article. Review.
-
Xeroderma pigmentosum-variant patients from America, Europe, and Asia.J Invest Dermatol. 2008 Aug;128(8):2055-68. doi: 10.1038/jid.2008.48. Epub 2008 Mar 27. J Invest Dermatol. 2008. PMID: 18368133 Free PMC article.
References
-
- Arlett C.F., Harcourt,S.A. and Broughton,B.C. (1975) The influence of caffeine on cell survival in excision-proficient and excision-deficient xeroderma pigmentosum and normal human cell strains following ultraviolet light irradiation. Mutat. Res., 33, 341–346. - PubMed
-
- Broomfield S., Hryciw,T. and Xiao,W. (2001) DNA postreplication repair and mutagenesis in Saccharomyces cerevisiae. Mutat. Res., 486, 167–184. - PubMed
-
- Cleaver J.E. et al. (1999) Increased ultraviolet sensitivity and chromosomal instability related to p53 function in the xeroderma pigmentosum variant. Cancer Res., 59, 1102–1108. - PubMed
-
- Friedberg E.C., Wagner,R. and Radman,M. (2002) Specialized DNA polymerases, cellular survival and the genesis of mutations. Science, 296, 1627–1630. - PubMed
-
- Goodman M.F. (2002) Error-prone repair DNA polymerases in prokaryotes and eukaryotes. Annu. Rev. Biochem., 71, 17–50. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

