Cellular mobile genetic elements in the regulatory region of the pneumotropic mouse polyomavirus genome: structure and function in viral gene expression and DNA replication
- PMID: 12610123
- PMCID: PMC149493
- DOI: 10.1128/jvi.77.6.3477-3486.2003
Cellular mobile genetic elements in the regulatory region of the pneumotropic mouse polyomavirus genome: structure and function in viral gene expression and DNA replication
Abstract
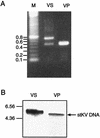
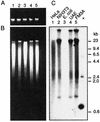
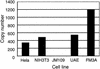
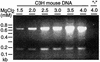
DNA from the murine pneumotropic virus was extracted from virus in lung tissue of infected mice, and the regulatory region of the genome was amplified by PCR. The regulatory region of individual plasmid cloned DNA molecules appeared to have heterogeneous enhancer segments, whereas the protein-coding part of the genome had a uniform length. Nucleotide sequence analysis revealed that the majority of the DNA molecules had a structure differing from the standard type. A 220-bp insertion at nucleotide position 142 with a concomitant deletion of nucleotides 143 to 148 was prominent. There were two variants of the 220-bp insertion, differing at two nucleotide positions at one of the termini. Other DNA molecules had complete or partial deletions of these structures and surrounding sequences in the viral enhancer. However, the end of the insertion at nucleotide 142 was frequently preserved. The viral early and late promoter activity of the variant regulatory regions was tested in a luciferase reporter assay by using transfected NIH 3T3 cells. In relation to the standard-type DNA, all variants, including a G272T mutant, had much stronger late promoters. In contrast, the early promoter activity was influenced in a positive or negative direction by individual mutations. Also, the activity of the viral origin of DNA replication was affected by the sequence variation of the regulatory region, although the effects were smaller than for the late promoter. Analysis by Southern blotting and quantification using dot blots showed that approximately 10(3) copies of material related to the 220-bp insert in murine pneumotropic virus DNA was present in mouse and human DNA but not in Escherichia coli DNA. Moreover, analysis by PCR indicated that there were multiple copies in the mouse genome of sequences that were identical or closely related to the 220-bp viral DNA segment. These data together with the nucleotide sequence analysis suggest that the 220-bp insertion is related to a transposable element of a novel type.
Figures


Similar articles
-
Extension of JC virus host range to monkey cells by insertion of a simian virus 40 enhancer into the JC virus regulatory region.Virology. 1989 Jun;170(2):353-61. doi: 10.1016/0042-6822(89)90425-x. Virology. 1989. PMID: 2543122
-
Kilham polyomavirus: activation of gene expression and DNA replication in mouse fibroblast cells by an enhancer substitution.J Virol. 2001 Nov;75(21):10015-23. doi: 10.1128/JVI.75.21.10015-10023.2001. J Virol. 2001. PMID: 11581370 Free PMC article.
-
Analysis of polyomavirus enhancer-effect on DNA replication and early gene expression.J Mol Biol. 1991 Apr 5;218(3):479-83. doi: 10.1016/0022-2836(91)90690-8. J Mol Biol. 1991. PMID: 1850000
-
A model for the formation of the duplicated enhancers found in polyomavirus regulatory regions.Virology. 2020 Apr;543:27-33. doi: 10.1016/j.virol.2020.01.013. Epub 2020 Jan 29. Virology. 2020. PMID: 32056844 Review.
-
RNA processing in the polyoma virus life cycle.Front Biosci (Landmark Ed). 2009 Jun 1;14(13):4968-77. doi: 10.2741/3581. Front Biosci (Landmark Ed). 2009. PMID: 19482599 Free PMC article. Review.
Cited by
-
Rearrangement of simian virus 40 regulatory region is not required for induction of progressive multifocal leukoencephalopathy in immunosuppressed rhesus monkeys.J Virol. 2005 Feb;79(3):1361-6. doi: 10.1128/JVI.79.3.1361-1366.2005. J Virol. 2005. PMID: 15650162 Free PMC article.
-
Complete Genome Sequence of Murine Pneumotropic Virus (Polyomaviridae) Clone pKV(37-1).Genome Announc. 2016 May 19;4(3):e00404-16. doi: 10.1128/genomeA.00404-16. Genome Announc. 2016. PMID: 27198030 Free PMC article.
-
Variable Genome Sequences of the Murine Pneumotropic Virus (Polyomaviridae) Regulatory Region Isolated from an Infected Mouse Tissue Viral Suspension.Genome Announc. 2016 May 26;4(3):e00405-16. doi: 10.1128/genomeA.00405-16. Genome Announc. 2016. PMID: 27231357 Free PMC article.
-
Receptor-binding and oncogenic properties of polyoma viruses isolated from feral mice.PLoS Pathog. 2007 Dec;3(12):e179. doi: 10.1371/journal.ppat.0030179. PLoS Pathog. 2007. PMID: 18085820 Free PMC article.
References
-
- Bastaki, M., E. E. Nelli, P. Dell'Era, M. Rusnati, M. P. Molinari-Tosatti, S. Parolini, R. Auerbach, L. P. Ruco, L. Possati, and M. Presta. 1997. Basic fibroblast growth factor-induced angiogenic phenotype in mouse endothelium. A study of aortic and microvascular endothelial cell lines. Arterioscler. Thromb. Vasc. Biol. 17:454-464. - PubMed
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

