Protein motion from non-specific to specific DNA by three-dimensional routes aided by supercoiling
- PMID: 12628933
- PMCID: PMC151056
- DOI: 10.1093/emboj/cdg125
Protein motion from non-specific to specific DNA by three-dimensional routes aided by supercoiling
Abstract
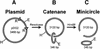
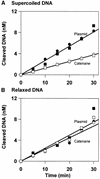
DNA-binding proteins are generally thought to locate their target sites by first associating with the DNA at random and then translocating to the specific site by one-dimensional (1D) diffusion along the DNA. We report here that non-specific DNA conveys proteins to their target sites just as well when held near the target by catenation as when co-linear with the target. Hence, contrary to the prevalent view, proteins move from random to specific sites primarily by three-dimensional (3D) rather than 1D pathways, by multiple dissociation/re-association events within a single DNA molecule. We also uncover a role for DNA supercoiling in target-site location. Proteins find their sites more readily in supercoiled than in relaxed DNA, again indicating 3D rather than 1D routes.
Figures

References
-
- Adzuma K. and Mizuuchi,K. (1989) Interaction of proteins located at a distance along DNA: mechanism of target immunity in the Mu DNA strand-transfer reaction. Cell, 57, 41–47. - PubMed
-
- Amouyal M. and Buc,H. (1987) Topological unwinding of strong and weak promoters by RNA polymerase. A comparison between the lac wild-type and UV5 sites of Escherichia coli. J. Mol. Biol., 195, 795–808. - PubMed
-
- Baldwin G.S., Sessions,R.B., Erskine,S.G. and Halford,S.E. (1999) DNA cleavage by the EcoRV restriction endonuclease: roles of divalent metal ions in specificity and catalysis. J. Mol. Biol., 288, 87–103. - PubMed
-
- Bates A.D. and Maxwell,A. (1993) DNA Topology. IRL Press, Oxford, UK.
-
- Berg H.C. (1993) Random Walks in Biology. Princeton University Press, Princeton, NJ.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

