Type III secretion systems and bacterial flagella: insights into their function from structural similarities
- PMID: 12631703
- PMCID: PMC152238
- DOI: 10.1073/pnas.0535335100
Type III secretion systems and bacterial flagella: insights into their function from structural similarities
Abstract
Type III secretion systems and bacterial flagella are broadly compared at the level of their genetic structure, morphology, regulation, and function, integrating structural information, to provide an overview of how they might function at a molecular level.
Figures
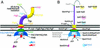
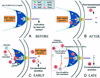


References
-
- Macnab R M. In: Escherichia coli and Salmonella: Cellular and Molecular Biology. Neidhardt F C, Curtis R III, Ingraham J L, Lin E C C, Low K B, Magasanik B, Reznikoff W S, Riley M, Schaechter M, Umbarger H E, editors. Vol. 1. Washington, DC: Am. Soc. Microbiol.; 1996. pp. 123–145.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

