Cloning and engineering of the cinnamycin biosynthetic gene cluster from Streptomyces cinnamoneus cinnamoneus DSM 40005
- PMID: 12642677
- PMCID: PMC153090
- DOI: 10.1073/pnas.0230516100
Cloning and engineering of the cinnamycin biosynthetic gene cluster from Streptomyces cinnamoneus cinnamoneus DSM 40005
Abstract
Lantibiotics are ribosomally synthesized oligopeptide antibiotics that contain lanthionine bridges derived by the posttranslational modification of amino acid residues. Here, we describe the cinnamycin biosynthetic gene cluster (cin) from Streptomyces cinnamoneus cinnamoneus DSM 40005, the first, to our knowledge, lantibiotic gene cluster from a high G+C bacterium to be cloned and sequenced. The cin cluster contains many genes not found in lantibiotic clusters from low G+C Gram-positive bacteria, including a Streptomyces antibiotic regulatory protein regulatory gene, and lacks others found in such clusters, such as a LanT-type transporter and a LanP-type protease. Transfer of the cin cluster to Streptomyces lividans resulted in heterologous production of cinnamycin. Furthermore, modification of the cinnamycin structural gene (cinA) led to production of two naturally occurring lantibiotics, duramycin and duramycin B, closely resembling cinnamycin, whereas attempts to make a more widely diverged derivative, duramycin C, failed to generate biologically active material. These results provide a basis for future attempts to construct extensive libraries of cinnamycin variants.
Figures



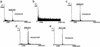
References
-
- Sahl H-G, Bierbaum B. Annu Rev Microbiol. 1998;52:41–79. - PubMed
-
- Jung G. In: Nisin and Novel Lantibiotics. Jung G, Sahl H A, editors. Leiden, The Netherlands: ESCOM; 1991. pp. 1–34.
-
- Brotz H, Josten M, Wiedemann I, Schneider U, Gotz F, Bierbaum G, Sahl H G. Mol Microbiol. 1998;30:317–327. - PubMed
-
- Witt D, Stackebrandt E. Syst Appl Microbiol. 1990;13:361–371.
-
- Choung S Y, Kobayashi T, Takemoto K, Ishitsuka H, Inoue K. Biochim Biophys Acta. 1998;940:180–187. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

