Recruitment of terminal protein to the ends of Streptomyces linear plasmids and chromosomes by a novel telomere-binding protein essential for linear DNA replication
- PMID: 12651895
- PMCID: PMC196017
- DOI: 10.1101/gad.1060303
Recruitment of terminal protein to the ends of Streptomyces linear plasmids and chromosomes by a novel telomere-binding protein essential for linear DNA replication
Abstract
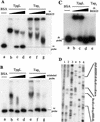
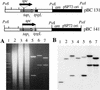
Bidirectional replication of Streptomyces linear plasmids and chromosomes from a central origin produces unpaired 3'-leading-strand overhangs at the telomeres of replication intermediates. Filling in of these overhangs leaves a terminal protein attached covalently to the 5' DNA ends of mature replicons. We report here the essential role of a novel 80-kD DNA-binding protein (telomere-associated protein, Tap) in this process. Biochemical studies, yeast two-hybrid analysis, and immunoprecipitation/immunodepletion experiments indicate that Tap binds tightly to specific sequences in 3' overhangs and also interacts with Tpg, bringing Tpg to telomere termini. Using DNA microarrays to analyze the chromosomes of tap mutant bacteria, we demonstrate that survivors of Tap ablation undergo telomere deletion, chromosome circularization, and amplification of subtelomeric DNA. Microarray-based chromosome mapping at single-ORF resolution revealed common endpoints for independent deletions, identified amplified chromosomal ORFs adjacent to these endpoints, and quantified the copy number of these ORFs. Sequence analysis confirmed chromosome circularization and revealed the insertion of adventitious DNA between joined chromosome ends. Our results show that Tap is required for linear DNA replication in Streptomyces and suggest that it functions to recruit and position Tpg at the telomeres of replication intermediates. They also identify hotspots for the telomeric deletions and subtelomeric DNA amplifications that accompany chromosome circularization.
Figures

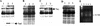



References
-
- Baumann P, Cech TR. Pot1, the putative telomere end-binding protein in fission yeast and humans. Science. 2001;292:1171–1175. - PubMed
-
- Behr MA, Wilson MA, Gill WP, Salamon H, Schoolnik GK, Rane S, Small PM. Comparative genomics of BCG vaccines by whole-genome DNA microarray. Science. 1999;284:1520–1523. - PubMed
-
- Bentley SD, Chater KF, Cerdeno-Tarraga AM, Challis GL, Thomson NR, James KD, Harris DE, Quail MA, Kieser H, Harper D, et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2) Nature. 2002;417:141–147. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
