Activities of atazanavir (BMS-232632) against a large panel of human immunodeficiency virus type 1 clinical isolates resistant to one or more approved protease inhibitors
- PMID: 12654666
- PMCID: PMC152527
- DOI: 10.1128/AAC.47.4.1324-1333.2003
Activities of atazanavir (BMS-232632) against a large panel of human immunodeficiency virus type 1 clinical isolates resistant to one or more approved protease inhibitors
Abstract
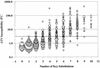
To evaluate the cross-resistance profile of the human immunodeficiency virus type 1 protease inhibitor (PI) atazanavir (BMS-232632), a panel of 551 clinical isolates exhibiting a wide array of PI resistance profiles and a variety of genotypic patterns were assayed for susceptibility to atazanavir and six other PIs: amprenavir, indinavir, lopinavir, nelfinavir, ritonavir, and saquinavir. In general, reductions in atazanavir susceptibility in vitro required several amino acid changes and were relatively modest in degree, and susceptibility was retained among isolates resistant to one or two of the currently approved PIs. There was a clear trend toward loss of susceptibility to atazanavir, as isolates exhibited increasing levels of cross-resistance to multiple PIs. Atazanavir appeared to have a distinct resistance profile relative to each of the other six PIs tested based on susceptibility comparisons against this panel of resistant isolates. Analysis of the genotypic profiles of 943 PI-susceptible and -resistant clinical isolates identified a strong correlation between the presence of amino acid changes at specific residues (10I/V/F, 20R/M/I, 24I, 33I/F/V, 36I/L/V, 46I/L, 48V, 54V/L, 63P, 71V/T/I, 73C/S/T/A, 82A/F/S/T, 84V, and 90M) and decreased susceptibility to atazanavir. While no single substitution or combination of substitutions was predictive of atazanavir resistance (change, >3.0-fold), the presence of at least five of these substitutions correlated strongly with loss of atazanavir susceptibility. Mutations associated with reduced susceptibility to each of the other six PIs were also determined.
Figures



References
-
- Angarano, G., and L. Monno. 2000. Genotype and phenotype resistance: an overview. J. Biol. Regul. Homeost. Agents 14:11-14. - PubMed
-
- Atkinson, B., J. Isaacson, M. Knowles, E. Mazabel, and A. K. Patick. 2000. Correlation between human immunodeficiency virus genotypic resistance and virologic response in patients receiving nelfinavir monotherapy or nelfinavir with lamivudine and zidovudine. J. Infect. Dis. 182:420-427. - PubMed
-
- Bally, F., R. Martinez, S. Peters, P. Sudre, and A. Telenti. 2000. Polymorphism of HIV type 1 gag p7/p1 and p1/p6 cleavage sites: clinical significance and implications for resistance to protease inhibitors. AIDS Res. Hum. Retroviruses 16:1209-1213. - PubMed
-
- Baxter, J. D., D. L. Mayers, D. N. Wentworth, J. D. Neaton, M. L. Hoover, M. A. Winters, S. B. Mannheimer, M. A. Thompson, D. I. Abrams, B. J. Brizz, J. P. Ioannidis, and T. C. Merigan. 2000. A randomized study of antiretroviral management based on plasma genotypic antiretroviral resistance testing in patients failing therapy. CPCRA 046 Study Team for the Terry Beirn Community Programs for Clinical Research on AIDS. AIDS 14:F83-93. - PubMed
-
- Boden, D., A. Hurley, L. Zhang, Y. Cao, Y. Guo, E. Jones, J. Tsay, J. Ip, C. Farthing, K. Limoli, N. Parkin, and M. Markowitz. 1999. HIV-1 drug resistance in newly infected individuals. JAMA 282:1135-1141. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

