Organization of the G protein-coupled receptors rhodopsin and opsin in native membranes
- PMID: 12663652
- PMCID: PMC1360145
- DOI: 10.1074/jbc.M302536200
Organization of the G protein-coupled receptors rhodopsin and opsin in native membranes
Abstract
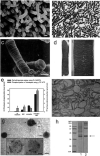
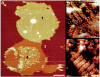
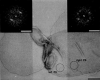
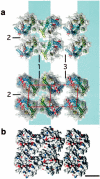
G protein-coupled receptors (GPCRs), which constitute the largest and structurally best conserved family of signaling molecules, are involved in virtually all physiological processes. Crystal structures are available only for the detergent-solubilized light receptor rhodopsin. In addition, this receptor is the only GPCR for which the presumed higher order oligomeric state in native membranes has been demonstrated (Fotiadis, D., Liang, Y., Filipek, S., Saperstein, D. A., Engel, A., and Palczewski, K. (2003) Nature 421, 127-128). Here, we have determined by atomic force microscopy the organization of rhodopsin in native membranes obtained from wild-type mouse photoreceptors and opsin isolated from photoreceptors of Rpe65-/- mutant mice, which do not produce the chromophore 11-cis-retinal. The higher order organization of rhodopsin was present irrespective of the support on which the membranes were adsorbed for imaging. Rhodopsin and opsin form structural dimers that are organized in paracrystalline arrays. The intradimeric contact is likely to involve helices IV and V, whereas contacts mainly between helices I and II and the cytoplasmic loop connecting helices V and VI facilitate the formation of rhodopsin dimer rows. Contacts between rows are on the extracellular side and involve helix I. This is the first semi-empirical model of a higher order structure of a GPCR in native membranes, and it has profound implications for the understanding of how this receptor interacts with partner proteins.
Figures


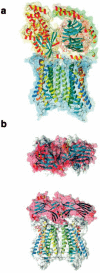
References
-
- Gehring WJ. Int. J. Dev. Biol. 2002;46:65–73. - PubMed
-
- Stryer L. Biopolymers. 1985;24:29–47. - PubMed
-
- Pierce KL, Premont RT, Lefkowitz RJ. Nat. Rev. Mol. Cell. Biol. 2002;3:639–650. - PubMed
-
- Palczewski K, Kumasaka T, Hori T, Behnke CA, Motoshima H, Fox BA, Le Trong I, Teller DC, Okada T, Stenkamp RE, Yamamoto M, Miyano M. Science. 2000;289:739–745. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

