The RUB/Nedd8 conjugation pathway is required for early development in Arabidopsis
- PMID: 12682009
- PMCID: PMC154480
- DOI: 10.1093/emboj/cdg190
The RUB/Nedd8 conjugation pathway is required for early development in Arabidopsis
Abstract
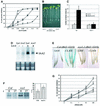
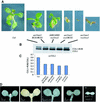
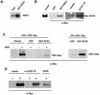
The related-to-ubiquitin (RUB) protein is post-translationally conjugated to the cullin subunit of the SCF (SKP1, Cullin, F-box) class of ubiquitin protein ligases. Although the precise biochemical function of RUB modification is unclear, studies indicate that the modification is important for SCF function. In Arabidopsis, RUB modification of CUL1 is required for normal function of SCF(TIR1), an E3 required for response to the plant hormone auxin. In this report we show that an Arabidopsis protein called RCE1 functions as a RUB-conjugating enzyme in vivo. A mutation in the RCE1 gene results in a phenotype like that of the axr1 mutant. Most strikingly, plants deficient in both RCE1 and AXR1 have an embryonic phenotype similar to mp and bdl mutants, previously shown to be deficient in auxin signaling. Based on these results, we suggest that the RUB-conjugation pathway is required for auxin-dependent pattern formation in the developing embryo. In addition, we show that RCE1 interacts directly with the RING protein RBX1 and is present in a stable complex with SCF. We propose that RBX1 functions as an E3 for RUB modification of CUL1.
Figures


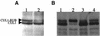
Similar articles
-
Role of the Arabidopsis RING-H2 protein RBX1 in RUB modification and SCF function.Plant Cell. 2002 Sep;14(9):2137-44. doi: 10.1105/tpc.003178. Plant Cell. 2002. PMID: 12215511 Free PMC article.
-
A new CULLIN 1 mutant has altered responses to hormones and light in Arabidopsis.Plant Physiol. 2007 Feb;143(2):684-96. doi: 10.1104/pp.106.091439. Epub 2006 Dec 8. Plant Physiol. 2007. PMID: 17158585 Free PMC article.
-
AXL and AXR1 have redundant functions in RUB conjugation and growth and development in Arabidopsis.Plant J. 2007 Oct;52(1):114-23. doi: 10.1111/j.1365-313X.2007.03211.x. Epub 2007 Jul 26. Plant J. 2007. PMID: 17655650
-
Jasmonate and auxin perception: how plants keep F-boxes in check.J Exp Bot. 2019 Jul 5;70(13):3401-3414. doi: 10.1093/jxb/erz272. J Exp Bot. 2019. PMID: 31173086 Review.
-
Regulation of cullin RING ligases.Annu Rev Plant Biol. 2008;59:467-89. doi: 10.1146/annurev.arplant.58.032806.104011. Annu Rev Plant Biol. 2008. PMID: 18444905 Review.
Cited by
-
Role of the MPN subunits in COP9 signalosome assembly and activity, and their regulatory interaction with Arabidopsis Cullin3-based E3 ligases.Plant Cell. 2007 Feb;19(2):564-81. doi: 10.1105/tpc.106.047571. Epub 2007 Feb 16. Plant Cell. 2007. PMID: 17307927 Free PMC article.
-
Mutation of E1-CONJUGATING ENZYME-RELATED1 decreases RELATED TO UBIQUITIN conjugation and alters auxin response and development.Plant Physiol. 2007 Jun;144(2):976-87. doi: 10.1104/pp.107.100404. Epub 2007 Apr 20. Plant Physiol. 2007. PMID: 17449645 Free PMC article.
-
DENEDDYLASE1 Protein Counters Automodification of Neddylating Enzymes to Maintain NEDD8 Protein Homeostasis in Arabidopsis.J Biol Chem. 2017 Mar 3;292(9):3854-3865. doi: 10.1074/jbc.M116.767103. Epub 2017 Jan 17. J Biol Chem. 2017. PMID: 28096463 Free PMC article.
-
AtCAND1, a HEAT-repeat protein that participates in auxin signaling in Arabidopsis.Plant Physiol. 2004 Jun;135(2):1020-6. doi: 10.1104/pp.104.044495. Epub 2004 Jun 4. Plant Physiol. 2004. PMID: 15181201 Free PMC article.
-
SPLICING FACTOR1 Is Important in Chloroplast Development under Cold Stress.Plant Physiol. 2020 Oct;184(2):973-987. doi: 10.1104/pp.20.00706. Epub 2020 Jul 30. Plant Physiol. 2020. PMID: 32732348 Free PMC article.
References
-
- Berleth T. and Jurgens,G. (1993) The role of the monopteros gene in organising the basal body region of the Arabidopsis embryo. Development, 118, 575–587.
-
- Berleth T., Mattsson,J. and Hardtke,C.S. (2000) Vascular continuity and auxin signals. Trends Plant Sci., 5, 387–393. - PubMed
-
- del Pozo J.C., Timpte,C., Tan,S., Callis,J. and Estelle,M. (1998) The ubiquitin-related protein RUB1 and auxin response in Arabidopsis. Science, 280, 1760–1763. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

