FGFR1 is independently required in both developing mid- and hindbrain for sustained response to isthmic signals
- PMID: 12682014
- PMCID: PMC154461
- DOI: 10.1093/emboj/cdg169
FGFR1 is independently required in both developing mid- and hindbrain for sustained response to isthmic signals
Abstract
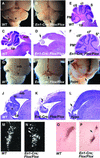
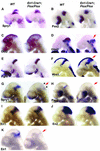
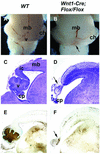
Fibroblast growth factors (FGFs) are signaling molecules of the isthmic organizer, which regulates development of the midbrain and cerebellum. Tissue-specific inactivation of one of the FGF receptor (FGFR) genes, Fgfr1, in the midbrain and rhombomere 1 of the hindbrain of mouse embryos results in deletion of the inferior colliculi in the posterior midbrain and vermis of the cerebellum. Analyses of both midbrain-hindbrain and midbrain-specific Fgfr1 mutants suggest that after establishment of the isthmic organizer, FGFR1 is needed for continued response to the isthmic signals, and that it has direct functions on both sides of the organizer. In addition, FGFR1 appears to modify cell adhesion properties critical for maintaining a coherent organizing center. This may be achieved by regulating expression of specific cell-adhesion molecules at the midbrain-hindbrain border.
Figures




Similar articles
-
Signaling specificities of fibroblast growth factor receptors in early Xenopus embryo.J Cell Sci. 2000 Aug;113 ( Pt 16):2865-75. doi: 10.1242/jcs.113.16.2865. J Cell Sci. 2000. PMID: 10910771
-
FGF regulated gene-expression and neuronal differentiation in the developing midbrain-hindbrain region.Dev Biol. 2006 Sep 1;297(1):141-57. doi: 10.1016/j.ydbio.2006.05.002. Epub 2006 May 13. Dev Biol. 2006. PMID: 16782087
-
Fgfr1-dependent boundary cells between developing mid- and hindbrain.Dev Biol. 2005 Feb 15;278(2):428-39. doi: 10.1016/j.ydbio.2004.11.024. Dev Biol. 2005. PMID: 15680361
-
FGF signalling pathways in development of the midbrain and anterior hindbrain.J Neurochem. 2007 Jun;101(5):1185-93. doi: 10.1111/j.1471-4159.2007.04463.x. Epub 2007 Feb 26. J Neurochem. 2007. PMID: 17326764 Review.
-
Neural plate patterning: upstream and downstream of the isthmic organizer.Nat Rev Neurosci. 2001 Feb;2(2):99-108. doi: 10.1038/35053516. Nat Rev Neurosci. 2001. PMID: 11253000 Review.
Cited by
-
A Versatile Gene Delivery System for Efficient and Tumor Specific Gene Manipulation in vivo.Discoveries (Craiova). 2016 Apr-Jun;4(2):e58. doi: 10.15190/d.2016.5. Epub 2016 Jun 22. Discoveries (Craiova). 2016. PMID: 27376150 Free PMC article.
-
FGFs: Neurodevelopment's Jack-of-all-Trades - How Do They Do it?Front Neurosci. 2011 Dec 5;5:133. doi: 10.3389/fnins.2011.00133. eCollection 2011. Front Neurosci. 2011. PMID: 22164131 Free PMC article.
-
Fgfr1 conditional-knockout in neural crest cells induces heterotopic chondrogenesis and osteogenesis in mouse frontal bones.Med Mol Morphol. 2019 Sep;52(3):156-163. doi: 10.1007/s00795-018-0213-z. Epub 2018 Nov 29. Med Mol Morphol. 2019. PMID: 30499042
-
Receptor tyrosine kinase (RTK) signalling in the control of neural stem and progenitor cell (NSPC) development.Mol Neurobiol. 2014 Feb;49(1):440-71. doi: 10.1007/s12035-013-8532-5. Epub 2013 Aug 28. Mol Neurobiol. 2014. PMID: 23982746 Review.
-
Sprouty genes control diastema tooth development via bidirectional antagonism of epithelial-mesenchymal FGF signaling.Dev Cell. 2006 Aug;11(2):181-90. doi: 10.1016/j.devcel.2006.05.014. Dev Cell. 2006. PMID: 16890158 Free PMC article.
References
-
- Acampora D., Avantaggiato,V., Tuorto,F. and Simeone,A. (1997) Genetic control of brain morphogenesis through Otx gene dosage requirement. Development, 124, 3639–3650. - PubMed
-
- Ang S.L. and Rossant,J. (1993) Anterior mesendoderm induces mouse Engrailed genes in explant cultures. Development, 118, 139–149. - PubMed
-
- Bally-Cuif L., Cholley,B. and Wassef,M. (1995) Involvement of Wnt-1 in the formation of the mes/metencephalic boundary. Mech. Dev., 53, 23–34. - PubMed
-
- Blair S.S. and Ralston,A. (1997) Smoothened-mediated Hedgehog signalling is required for the maintenance of the anterior–posterior lineage restriction in the developing wing of Drosophila. Development, 124, 4053–4063. - PubMed
-
- Broccoli V., Boncinelli,E. and Wurst,W. (1999) The caudal limit of Otx2 expression positions the isthmic organizer. Nature, 401, 164–168. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

