Haemophilus influenzae carriage in children attending French day care centers: a molecular epidemiological study
- PMID: 12682158
- PMCID: PMC153885
- DOI: 10.1128/JCM.41.4.1664-1672.2003
Haemophilus influenzae carriage in children attending French day care centers: a molecular epidemiological study
Abstract
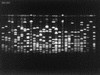
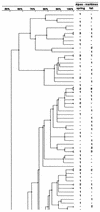
The nasopharyngeal Haemophilus influenzae flora of healthy children under the age of 3 years attending day care centers in three distinct French geographic areas was analyzed by sampling during two periods, spring 1999 (May and June) and fall 1999 (November and December). The average carrier rate among 1,683 children was 40.9%. The prevalence of capsulated H. influenzae carriers was 0.4% for type f and 0.6% for type e. No type b strains were found among these children, of whom 98.5% had received one or more doses of anti-Haemophilus b vaccine. Among the strains, 44.5% were TEM-type beta-lactamase producers and nine (1.3%) were beta-lactamase-negative ampicillin-resistant strains. Pulsed-field gel electrophoresis restriction patterns showed a large diversity with 366 SmaI patterns from 663 strains. Among the strains isolated during a given period, 33% were isolated simultaneously in more than one area. In each area, depending on the sampling period, 68 to 72% of the strains had new pulsotypes and persistence of 28 to 32% of the strains was noted. For the 297 beta-lactamase-producing strains, 194 patterns were found. The genomic diversity of these strains was comparable to that of the whole set of strains and does not suggest a clonal diffusion. Among the beta-lactamase-producing strains isolated in November and December, depending on the area, 66 to 73% had new pulsotypes with persistence of only 27 to 33% of the strains. In any given geographic area, colonization by H. influenzae appears to be a dynamic process involving a high degree of genomic heterogeneity among the noncapsulated colonizing strains.
Figures






References
-
- Bernstein, J. M., D. Dryja, N. Yuskiw, and T. F. Murphy. 1997. Analysis of isolates recovered from multiple sites of the nasopharynx of children colonized by nontypeable Haemophilus influenzae. Eur. J. Clin. Microbiol. Infect. Dis. 16:750-753. - PubMed
-
- Bou, R., À. Domínguez, D. Fontanals, I. Sanfeliu, I. Pons, J. Renau, V. Pineda, E. Lobera, C. Latorre, M. Majó, L. Salleras, and the Working Group on invasive disease caused by Haemophilus influenzae. 2000. Prevalence of Haemophilus influenzae pharyngeal carriers in the school population of Catalonia. Eur. J. Epidemiol. 16:521-526. - PubMed
-
- Comité de l'Antibiogramme de la Société Française de Microbiologie. Communiqué 2002. [Online.] http://www.sfm.asso.fr/Sect4/com2002.pdf.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

