Characterization of the DNA-binding properties of the origin-binding domain of simian virus 40 large T antigen by fluorescence anisotropy
- PMID: 12692254
- PMCID: PMC153955
- DOI: 10.1128/jvi.77.9.5512-5518.2003
Characterization of the DNA-binding properties of the origin-binding domain of simian virus 40 large T antigen by fluorescence anisotropy
Abstract
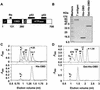
The affinity of the origin-binding domain (OBD) of simian virus 40 large T antigen for its cognate origin was measured at equilibrium using a DNA binding assay based on fluorescence anisotropy. At a near-physiological concentration of salt, the affinities of the OBD for site II and the core origin were 31 and 50 nM, respectively. Binding to any of the four 5'-GAGGC-3' binding sites in site II was only slightly weaker, between 57 and 150 nM. Although the OBD was shown previously to assemble as a dimer on two binding sites spaced by 7 bp, we found that increasing the distance between both binding sites by 1 to 3 bp had little effect on affinity. Similar results were obtained for full-length T antigen in absence of nucleotide. Addition of ADP-Mg, which promotes hexamerization of T antigen, greatly increased the affinity of full-length T antigen for the core origin and for nonspecific DNA. The implications of these findings for the assembly of T antigen at the origin and its transition to a non-specific DNA helicase are discussed.
Figures



Similar articles
-
Analysis of the costructure of the simian virus 40 T-antigen origin binding domain with site I reveals a correlation between GAGGC spacing and spiral assembly.J Virol. 2013 Mar;87(5):2923-34. doi: 10.1128/JVI.02549-12. Epub 2012 Dec 26. J Virol. 2013. PMID: 23269808 Free PMC article.
-
Quantitative analysis of the binding of simian virus 40 large T antigen to DNA.J Virol. 2007 Sep;81(17):9162-74. doi: 10.1128/JVI.00384-07. Epub 2007 Jun 27. J Virol. 2007. PMID: 17596312 Free PMC article.
-
T antigen origin-binding domain of simian virus 40: determinants of specific DNA binding.Biochemistry. 2004 Jun 8;43(22):6928-36. doi: 10.1021/bi030228+. Biochemistry. 2004. PMID: 15170330
-
Origin DNA-binding proteins.Curr Opin Struct Biol. 1998 Feb;8(1):49-53. doi: 10.1016/s0959-440x(98)80009-2. Curr Opin Struct Biol. 1998. PMID: 9519296 Review.
-
Interactions between DNA replication-related proteins and phospholipid vesicles in vitro.Chem Phys Lipids. 1994 Sep 6;73(1-2):223-30. doi: 10.1016/0009-3084(94)90183-x. Chem Phys Lipids. 1994. PMID: 8001183 Review.
Cited by
-
Analysis of the costructure of the simian virus 40 T-antigen origin binding domain with site I reveals a correlation between GAGGC spacing and spiral assembly.J Virol. 2013 Mar;87(5):2923-34. doi: 10.1128/JVI.02549-12. Epub 2012 Dec 26. J Virol. 2013. PMID: 23269808 Free PMC article.
-
Interactions required for binding of simian virus 40 T antigen to the viral origin and molecular modeling of initial assembly events.J Virol. 2004 Mar;78(6):2921-34. doi: 10.1128/jvi.78.6.2921-2934.2004. J Virol. 2004. PMID: 14990710 Free PMC article.
-
Origin DNA Melting-An Essential Process with Divergent Mechanisms.Genes (Basel). 2017 Jan 11;8(1):26. doi: 10.3390/genes8010026. Genes (Basel). 2017. PMID: 28085061 Free PMC article. Review.
-
Using fluorophore-labeled oligonucleotides to measure affinities of protein-DNA interactions.Methods Enzymol. 2008;450:253-72. doi: 10.1016/S0076-6879(08)03412-5. Methods Enzymol. 2008. PMID: 19152864 Free PMC article. Review.
-
Insights into hRPA32 C-terminal domain--mediated assembly of the simian virus 40 replisome.Nat Struct Mol Biol. 2005 Apr;12(4):332-9. doi: 10.1038/nsmb916. Epub 2005 Mar 27. Nat Struct Mol Biol. 2005. PMID: 15793585 Free PMC article.
References
-
- Bullock, P. A. 1997. The initiation of simian virus 40 DNA replication in vitro. Crit. Rev. Biochem. Mol. Biol. 32:503-568. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

