Binding to EGF receptor of a laminin-5 EGF-like fragment liberated during MMP-dependent mammary gland involution
- PMID: 12695504
- PMCID: PMC2172889
- DOI: 10.1083/jcb.200208145
Binding to EGF receptor of a laminin-5 EGF-like fragment liberated during MMP-dependent mammary gland involution
Abstract
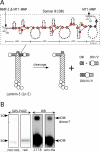
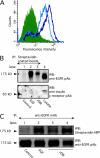
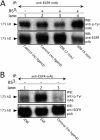
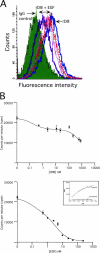
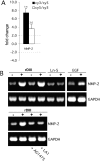
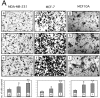
Extracellular matrix (ECM) fragments or cryptic sites unmasked by proteinases have been postulated to affect tissue remodeling and cancer progression. Therefore, the elucidation of their identities and functions is of great interest. Here, we show that matrix metalloproteinases (MMPs) generate a domain (DIII) from the ECM macromolecule laminin-5. Binding of a recombinant DIII fragment to epidermal growth factor receptor stimulates downstream signaling (mitogen-activated protein kinase), MMP-2 gene expression, and cell migration. Appearance of this cryptic ECM ligand in remodeling mammary gland coincides with MMP-mediated involution in wild-type mice, but not in tissue inhibitor of metalloproteinase 3 (TIMP-3)-deficient mice, supporting physiological regulation of DIII liberation. These findings indicate that ECM cues may operate via direct stimulation of receptor tyrosine kinases in tissue remodeling, and possibly cancer invasion.
Figures






Similar articles
-
The invasive phenotype in HMT-3522 cells requires increased EGF receptor signaling through both PI 3-kinase and ERK 1,2 pathways.Cell Commun Adhes. 2002 Mar-Apr;9(2):87-102. doi: 10.1080/15419060214147. Cell Commun Adhes. 2002. PMID: 12487410
-
ERK1/2 and p38 MAP kinase control MMP-2, MT1-MMP, and TIMP action and affect cell migration: a comparison between mesothelioma and mesothelial cells.J Cell Physiol. 2006 May;207(2):540-52. doi: 10.1002/jcp.20605. J Cell Physiol. 2006. PMID: 16447244
-
3-phosphoinositide-dependent protein kinase-1 (PDK1) promotes invasion and activation of matrix metalloproteinases.BMC Cancer. 2006 Mar 21;6:77. doi: 10.1186/1471-2407-6-77. BMC Cancer. 2006. PMID: 16551362 Free PMC article.
-
[The role of matrix metalloproteinases in the pathogenesis of diabetes mellitus and progression of diabetes retinopathy].Postepy Hig Med Dosw (Online). 2008 Sep 3;62:442-50. Postepy Hig Med Dosw (Online). 2008. PMID: 18772849 Review. Polish.
-
Matrix metalloproteinases in remodeling of the normal and neoplastic mammary gland.J Mammary Gland Biol Neoplasia. 1998 Apr;3(2):177-89. doi: 10.1023/a:1018746923474. J Mammary Gland Biol Neoplasia. 1998. PMID: 10819526 Review.
Cited by
-
Complex Regulation of the Pericellular Proteolytic Microenvironment during Tumor Progression and Wound Repair: Functional Interactions between the Serine Protease and Matrix Metalloproteinase Cascades.Biochem Res Int. 2012;2012:454368. doi: 10.1155/2012/454368. Epub 2012 Feb 20. Biochem Res Int. 2012. PMID: 22454771 Free PMC article.
-
Up-regulation of angiopoietin-2, matrix metalloprotease-2, membrane type 1 metalloprotease, and laminin 5 gamma 2 correlates with the invasiveness of human glioma.Am J Pathol. 2005 Mar;166(3):877-90. doi: 10.1016/s0002-9440(10)62308-5. Am J Pathol. 2005. PMID: 15743799 Free PMC article.
-
Lysophosphatidic Acid Upregulates Laminin-332 Expression during A431 Cell Colony Dispersal.J Oncol. 2010;2010:107075. doi: 10.1155/2010/107075. Epub 2010 Aug 25. J Oncol. 2010. PMID: 20862207 Free PMC article.
-
Laminin-5 in epithelial tumour invasion.J Mol Histol. 2004 Mar;35(3):277-86. doi: 10.1023/b:hijo.0000032359.35698.fe. J Mol Histol. 2004. PMID: 15339047 Review.
-
A hypomorphic mutation in the mouse laminin alpha5 gene causes polycystic kidney disease.J Am Soc Nephrol. 2006 Jul;17(7):1913-22. doi: 10.1681/ASN.2005121298. Epub 2006 Jun 21. J Am Soc Nephrol. 2006. PMID: 16790509 Free PMC article.
References
-
- Belkin, A.M., and M.A. Stepp. 2000. Integrins as receptors for laminins. Microsc. Res. Tech. 51:280–301. - PubMed
-
- Bilban, M., L.K. Buehler, S. Head, G. Desoye, and V. Quaranta. 2002. Normalizing DNA microarray data. Curr. Issues. Mol. Biol. 4:57–64. - PubMed
-
- Borradori, L., and A. Sonnenberg. 1999. Structure and function of hemidesmosomes: more than simple adhesion complexes. J. Invest. Dermatol. 112:411–418. - PubMed
-
- Chen, Z., T.B. Gibson, F. Robinson, L. Silvestro, G. Pearson, B. Xu, A. Wright, C. Vanderbilt, and M.H. Cobb. 2001. MAP Kinases. Chem. Rev. 101:2449–2476. - PubMed
-
- Csordas, G., M. Santra, C.C. Reed, I. Eichstetter, D.J. McQuillan, D. Gross, M.A. Nugent, G. Hajnoczky, and R.V. Iozzo. 2000. Sustained down-regulation of the epidermal growth factor receptor by decorin. A mechanism for controlling tumor growth in vivo. J. Biol. Chem. 275:32879–32887. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

