Two Drosophila Ada2 homologues function in different multiprotein complexes
- PMID: 12697829
- PMCID: PMC153191
- DOI: 10.1128/MCB.23.9.3305-3319.2003
Two Drosophila Ada2 homologues function in different multiprotein complexes
Abstract
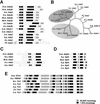
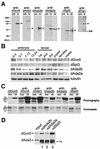
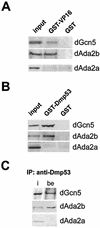
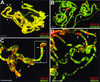
The reversible acetylation of the N-terminal tails of histones is crucial for transcription, DNA repair, and replication. The enzymatic reaction is catalyzed by large multiprotein complexes, of which the best characterized are the Gcn5-containing N-acetyltransferase (GNAT) complexes. GNAT complexes from yeast to humans share several conserved subunits, such as Ada2, Ada3, Spt3, and Tra1/TRRAP. We have characterized these factors in Drosophila and found that the flies have two distinct Ada2 variants (dAda2a and dAda2b). Using a combination of biochemical and cell biological approaches we demonstrate that only one of the two Drosophila Ada2 homologues, dAda2b, is a component of Spt-Ada-Gcn5-acetyltransferase (SAGA) complexes. The other Ada2 variant, dAda2a, can associate with dGcn5 but is not incorporated into dSAGA-type complexes. This is the first example of a complex-specific association of the Ada-type transcriptional adapter proteins with GNATs. In addition, dAda2a is part of Gcn5-independent complexes, which are concentrated at transcriptionally active regions on polytene chromosomes. This implicates novel functions for dAda2a in transcription. Humans and mice also possess two Ada2 variants with high homology to dAda2a and dAda2b, respectively. This suggests that the mammalian and fly homologues of the transcriptional adapter Ada2 form two functionally distinct subgroups with unique characteristics.
Figures





Similar articles
-
Two different Drosophila ADA2 homologues are present in distinct GCN5 histone acetyltransferase-containing complexes.Mol Cell Biol. 2003 Jan;23(1):306-21. doi: 10.1128/MCB.23.1.306-321.2003. Mol Cell Biol. 2003. PMID: 12482983 Free PMC article.
-
Host cell factor and an uncharacterized SANT domain protein are stable components of ATAC, a novel dAda2A/dGcn5-containing histone acetyltransferase complex in Drosophila.Mol Cell Biol. 2006 Feb;26(3):871-82. doi: 10.1128/MCB.26.3.871-882.2006. Mol Cell Biol. 2006. PMID: 16428443 Free PMC article.
-
Drosophila Ada2b is required for viability and normal histone H3 acetylation.Mol Cell Biol. 2004 Sep;24(18):8080-9. doi: 10.1128/MCB.24.18.8080-8089.2004. Mol Cell Biol. 2004. PMID: 15340070 Free PMC article.
-
The Gcn5 complexes in Drosophila as a model for metazoa.Biochim Biophys Acta Gene Regul Mech. 2021 Feb;1864(2):194610. doi: 10.1016/j.bbagrm.2020.194610. Epub 2020 Jul 28. Biochim Biophys Acta Gene Regul Mech. 2021. PMID: 32735945 Review.
-
The Ada2/Ada3/Gcn5/Sgf29 histone acetyltransferase module.Biochim Biophys Acta Gene Regul Mech. 2021 Feb;1864(2):194629. doi: 10.1016/j.bbagrm.2020.194629. Epub 2020 Sep 2. Biochim Biophys Acta Gene Regul Mech. 2021. PMID: 32890768 Free PMC article. Review.
Cited by
-
Ataxin-7 and Non-stop coordinate SCAR protein levels, subcellular localization, and actin cytoskeleton organization.Elife. 2019 Jul 26;8:e49677. doi: 10.7554/eLife.49677. Elife. 2019. PMID: 31348003 Free PMC article.
-
The Drosophila NURF remodelling and the ATAC histone acetylase complexes functionally interact and are required for global chromosome organization.EMBO Rep. 2008 Feb;9(2):187-92. doi: 10.1038/sj.embor.7401141. Epub 2007 Dec 14. EMBO Rep. 2008. PMID: 18084186 Free PMC article.
-
Histone H3 lysine 4 trimethylation regulates cotranscriptional H2A variant exchange by Tip60 complexes to maximize gene expression.Proc Natl Acad Sci U S A. 2014 Apr 1;111(13):4850-5. doi: 10.1073/pnas.1320337111. Epub 2014 Mar 17. Proc Natl Acad Sci U S A. 2014. PMID: 24639513 Free PMC article.
-
Histone acetylation, acetyltransferases, and ataxia--alteration of histone acetylation and chromatin dynamics is implicated in the pathogenesis of polyglutamine-expansion disorders.Adv Protein Chem Struct Biol. 2010;79:165-203. doi: 10.1016/S1876-1623(10)79005-2. Adv Protein Chem Struct Biol. 2010. PMID: 20621284 Free PMC article. Review.
-
Drosophila USP22/nonstop polarizes the actin cytoskeleton during collective border cell migration.J Cell Biol. 2021 Jul 5;220(7):e202007005. doi: 10.1083/jcb.202007005. Epub 2021 May 14. J Cell Biol. 2021. PMID: 33988679 Free PMC article.
References
-
- Aasland, R., A. F. Stewart, and T. Gibson. 1996. The SANT domain: a putative DNA-binding domain in the SWI-SNF and ADA complexes, the transcriptional co-repressor N-CoR and TFIIIB. Trends Biochem. Sci. 21:87-88. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Balasubramanian, R., M. G. Pray-Grant, W. Selleck, P. A. Grant, and S. Tan. 2002. Role of the Ada2 and Ada3 transcriptional coactivators in histone acetylation. J. Biol. Chem. 277:7989-7995. - PubMed
-
- Berger, S. L., B. Pina, N. Silverman, G. A. Marcus, J. Agapite, J. L. Regier, S. J. Triezenberg, and L. Guarente. 1992. Genetic isolation of ADA2: a potential transcriptional adaptor required for function of certain acidic activation domains. Cell 70:251-265. - PubMed
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
