Dimerization of MLH1 and PMS2 limits nuclear localization of MutLalpha
- PMID: 12697830
- PMCID: PMC153201
- DOI: 10.1128/MCB.23.9.3320-3328.2003
Dimerization of MLH1 and PMS2 limits nuclear localization of MutLalpha
Abstract
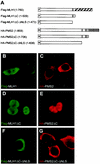
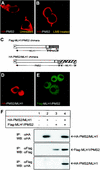
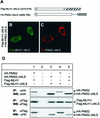
DNA mismatch repair maintains genomic stability by detecting and correcting mispaired DNA sequences and by signaling cell death when DNA repair fails. The mechanism by which mismatch repair coordinates DNA damage and repair with cell survival or death is not understood, but it suggests the need for regulation. Since the functions of mismatch repair are initiated in the nucleus, we asked whether nuclear transport of MLH1 and PMS2 is limiting for the nuclear localization of MutLalpha (the MLH1-PMS2 dimer). We found that MLH1 and PMS2 have functional nuclear localization signals (NLS) and nuclear export sequences, yet nuclear import depended on their C-terminal dimerization to form MutLalpha. Our studies are consistent with the idea that dimerization of MLH1 and PMS2 regulates nuclear import by unmasking the NLS. Limited nuclear localization of MutLalpha may thus represent a novel mechanism by which cells fine-tune mismatch repair functions. This mechanism may have implications in the pathogenesis of hereditary non-polyposis colon cancer.
Figures





Similar articles
-
Nuclear import of human MLH1, PMS2, and MutLalpha: redundancy is the key.Mol Carcinog. 2009 Aug;48(8):742-50. doi: 10.1002/mc.20514. Mol Carcinog. 2009. PMID: 19148896
-
Contributions by MutL homologues Mlh3 and Pms2 to DNA mismatch repair and tumor suppression in the mouse.Cancer Res. 2005 Oct 1;65(19):8662-70. doi: 10.1158/0008-5472.CAN-05-0742. Cancer Res. 2005. PMID: 16204034
-
Contribution of human mlh1 and pms2 ATPase activities to DNA mismatch repair.J Biol Chem. 2002 Jun 14;277(24):21801-9. doi: 10.1074/jbc.M111342200. Epub 2002 Mar 15. J Biol Chem. 2002. PMID: 11897781
-
[Homologs of MutS and MutL during mammalian meiosis].Med Sci (Paris). 2003 Jan;19(1):85-91. doi: 10.1051/medsci/200319185. Med Sci (Paris). 2003. PMID: 12836196 Review. French.
-
Mismatch repair pathway: molecules, functions, and role in colorectal carcinogenesis.Eur J Cancer Prev. 2014 Jul;23(4):246-57. doi: 10.1097/CEJ.0000000000000019. Eur J Cancer Prev. 2014. PMID: 24614649 Review.
Cited by
-
An intact Pms2 ATPase domain is not essential for male fertility.DNA Repair (Amst). 2016 Mar;39:46-51. doi: 10.1016/j.dnarep.2015.12.011. Epub 2015 Dec 29. DNA Repair (Amst). 2016. PMID: 26753533 Free PMC article.
-
Analysis of clinically relevant somatic mutations in high-risk head and neck cutaneous squamous cell carcinoma.Mod Pathol. 2018 Feb;31(2):275-287. doi: 10.1038/modpathol.2017.128. Epub 2017 Oct 6. Mod Pathol. 2018. PMID: 28984303
-
Structure of the MutL C-terminal domain: a model of intact MutL and its roles in mismatch repair.EMBO J. 2004 Oct 27;23(21):4134-45. doi: 10.1038/sj.emboj.7600412. Epub 2004 Oct 7. EMBO J. 2004. PMID: 15470502 Free PMC article.
-
Apoptotic function of human PMS2 compromised by the nonsynonymous single-nucleotide polymorphic variant R20Q.Proc Natl Acad Sci U S A. 2008 Sep 16;105(37):13993-8. doi: 10.1073/pnas.0806435105. Epub 2008 Sep 3. Proc Natl Acad Sci U S A. 2008. PMID: 18768816 Free PMC article.
-
Human PMS2 deficiency is associated with impaired immunoglobulin class switch recombination.J Exp Med. 2008 Oct 27;205(11):2465-72. doi: 10.1084/jem.20080789. Epub 2008 Sep 29. J Exp Med. 2008. PMID: 18824584 Free PMC article.
References
-
- Baker, S. M., C. E. Bronner, L. Zhang, A. W. Plug, M. Robatzek, G. Warren, E. A. Elliott, J. Yu, T. Ashley, N. Arnheim, R. A. Flavell, and R. M. Liskay. 1995. Male mice defective in the DNA mismatch repair gene PMS2 exhibit abnormal chromosome synapsis in meiosis. Cell 82:309-319. - PubMed
-
- Ban, C., and W. Yang. 1998. Crystal structure and ATPase activity of MutL: implications for DNA repair and mutagenesis. Cell 95:541-552. - PubMed
-
- Bellacosa, A. 2001. Functional interactions and signaling properties of mammalian DNA mismatch repair proteins. Cell Death Differ. 8:1076-1092. - PubMed
-
- Carmo-Fonseca, M. 2002. The contribution of nuclear compartmentalization to gene regulation. Cell 108:513-521. - PubMed
-
- Cascalho, M., J. Wong, C. Steinberg, and M. Wabl. 1998. Mismatch repair co-opted by hypermutation. Science 279:1207-1210. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
