Regulation of the bone-specific osteocalcin gene by p300 requires Runx2/Cbfa1 and the vitamin D3 receptor but not p300 intrinsic histone acetyltransferase activity
- PMID: 12697832
- PMCID: PMC153185
- DOI: 10.1128/MCB.23.9.3339-3351.2003
Regulation of the bone-specific osteocalcin gene by p300 requires Runx2/Cbfa1 and the vitamin D3 receptor but not p300 intrinsic histone acetyltransferase activity
Abstract
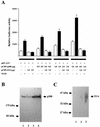
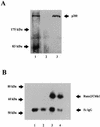
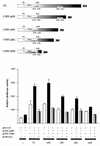
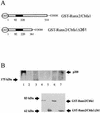
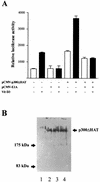
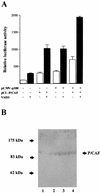
p300 is a multifunctional transcriptional coactivator that serves as an adapter for several transcription factors including nuclear steroid hormone receptors. p300 possesses an intrinsic histone acetyltransferase (HAT) activity that may be critical for promoting steroid-dependent transcriptional activation. In osteoblastic cells, transcription of the bone-specific osteocalcin (OC) gene is principally regulated by the Runx2/Cbfa1 transcription factor and is stimulated in response to vitamin D(3) via the vitamin D(3) receptor complex. Therefore, we addressed p300 control of basal and vitamin D(3)-enhanced activity of the OC promoter. We find that transient overexpression of p300 results in a significant dose-dependent increase of both basal and vitamin D(3)-stimulated OC gene activity. This stimulatory effect requires intact Runx2/Cbfa1 binding sites and the vitamin D-responsive element. In addition, by coimmunoprecipitation, we show that the endogenous Runx2/Cbfa1 and p300 proteins are components of the same complexes within osteoblastic cells under physiological concentrations. We also demonstrate by chromatin immunoprecipitation assays that p300, Runx2/Cbfa1, and 1alpha,25-dihydroxyvitamin D(3) receptor interact with the OC promoter in intact osteoblastic cells expressing this gene. The effect of p300 on the OC promoter is independent of its intrinsic HAT activity, as a HAT-deficient p300 mutant protein up-regulates expression and cooperates with P/CAF to the same extent as the wild-type p300. On the basis of these results, we propose that p300 interacts with key transcriptional regulators of the OC gene and bridges distal and proximal OC promoter sequences to facilitate responsiveness to vitamin D(3).
Figures




Similar articles
-
Bone-specific transcription factor Runx2 interacts with the 1alpha,25-dihydroxyvitamin D3 receptor to up-regulate rat osteocalcin gene expression in osteoblastic cells.Mol Cell Biol. 2004 Oct;24(20):8847-61. doi: 10.1128/MCB.24.20.8847-8861.2004. Mol Cell Biol. 2004. PMID: 15456860 Free PMC article.
-
1alpha,25-dihydroxy vitamin D3-enhanced expression of the osteocalcin gene involves increased promoter occupancy of basal transcription regulators and gradual recruitment of the 1alpha,25-dihydroxy vitamin D3 receptor-SRC-1 coactivator complex.J Cell Physiol. 2008 Mar;214(3):740-9. doi: 10.1002/jcp.21267. J Cell Physiol. 2008. PMID: 17786964
-
The Runx2 transcription factor plays a key role in the 1alpha,25-dihydroxy Vitamin D3-dependent upregulation of the rat osteocalcin (OC) gene expression in osteoblastic cells.J Steroid Biochem Mol Biol. 2004 May;89-90(1-5):269-71. doi: 10.1016/j.jsbmb.2004.03.076. J Steroid Biochem Mol Biol. 2004. PMID: 15225783 Review.
-
Identification of potential modifiers of Runx2/Cbfa1 activity in C2C12 cells in response to bone morphogenetic protein-7.Cells Tissues Organs. 2004;176(1-3):28-40. doi: 10.1159/000075025. Cells Tissues Organs. 2004. PMID: 14745233
-
Runx2/Cbfa1 functions: diverse regulation of gene transcription by chromatin remodeling and co-regulatory protein interactions.Connect Tissue Res. 2003;44 Suppl 1:141-8. Connect Tissue Res. 2003. PMID: 12952188 Review.
Cited by
-
Dlx3 transcriptional regulation of osteoblast differentiation: temporal recruitment of Msx2, Dlx3, and Dlx5 homeodomain proteins to chromatin of the osteocalcin gene.Mol Cell Biol. 2004 Oct;24(20):9248-61. doi: 10.1128/MCB.24.20.9248-9261.2004. Mol Cell Biol. 2004. PMID: 15456894 Free PMC article.
-
The Role of RUNX2 in Osteosarcoma Oncogenesis.Sarcoma. 2011;2011:282745. doi: 10.1155/2011/282745. Epub 2010 Dec 9. Sarcoma. 2011. PMID: 21197465 Free PMC article.
-
Protein arginine methyltransferases PRMT1, PRMT4/CARM1 and PRMT5 have distinct functions in control of osteoblast differentiation.Bone Rep. 2023 Jul 25;19:101704. doi: 10.1016/j.bonr.2023.101704. eCollection 2023 Dec. Bone Rep. 2023. PMID: 37593409 Free PMC article.
-
Effects of dexamethasone, ascorbic acid and β-glycerophosphate on the osteogenic differentiation of stem cells in vitro.Stem Cell Res Ther. 2013;4(5):117. doi: 10.1186/scrt328. Stem Cell Res Ther. 2013. PMID: 24073831 Free PMC article. Review.
-
Vitamin D receptor signaling mechanisms: integrated actions of a well-defined transcription factor.Steroids. 2013 Feb;78(2):127-36. doi: 10.1016/j.steroids.2012.10.019. Epub 2012 Nov 23. Steroids. 2013. PMID: 23178257 Free PMC article. Review.
References
-
- Aslam, F., L. McCabe, B. Frenkel, A. van Wijnen, G. S. Stein, J. B. Lian, and J. Stein. 1997. AP-1 and vitamin D receptor (VDR) signaling pathways converge at the rat osteocalcin VDR element: requirement for the internal activating protein-1 site for vitamin D-mediated trans-activation. Endocrinology 140:63-70. - PubMed
-
- Banerjee, C., L. McCabe, J. Choi, S. Hiebert, J. Stein, G. Stein, and J. Lian. 1997. Runt homology domain proteins in osteoblast differentiation: AML3/CBFA1 is a major component of a bone-specific complex. J. Cell. Biochem. 66:1-8. - PubMed
-
- Chakravarti, D., V. J. La Morte, M. C. Nelson, T. Nakajima, I. G. Schulman, H. Juguilon, M. Montminy, and R. M. Evans. 1996. Role of CBP/p300 in nuclear receptor signalling. Nature (London) 383:9-13. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
