A 1-megadalton ESC/E(Z) complex from Drosophila that contains polycomblike and RPD3
- PMID: 12697833
- PMCID: PMC153183
- DOI: 10.1128/MCB.23.9.3352-3362.2003
A 1-megadalton ESC/E(Z) complex from Drosophila that contains polycomblike and RPD3
Abstract
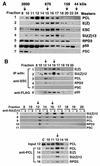
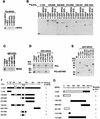
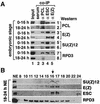
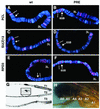
Polycomb group (PcG) proteins are required to maintain stable repression of the homeotic genes and others throughout development. The PcG proteins ESC and E(Z) are present in a prominent 600-kDa complex as well as in a number of higher-molecular-mass complexes. Here we identify and characterize a 1-MDa ESC/E(Z) complex that is distinguished from the 600-kDa complex by the presence of the PcG protein Polycomblike (PCL) and the histone deacetylase RPD3. In addition, the 1-MDa complex shares with the 600-kDa complex the histone binding protein p55 and the PcG protein SU(Z)12. Coimmunoprecipitation assays performed on embryo extracts and gel filtration column fractions indicate that, during embryogenesis E(Z), SU(Z)12, and p55 are present in all ESC complexes, while PCL and RPD3 are associated with ESC, E(Z), SU(Z)12, and p55 only in the 1-MDa complex. Glutathione transferase pulldown assays demonstrate that RPD3 binds directly to PCL via the conserved PHD fingers of PCL and the N terminus of RPD3. PCL and E(Z) colocalize virtually completely on polytene chromosomes and are associated with a subset of RPD3 sites. As previously shown for E(Z) and RPD3, PCL and SU(Z)12 are also recruited to the insertion site of a minimal Ubx Polycomb response element transgene in vivo. Consistent with these biochemical and cytological results, Rpd3 mutations enhance the phenotypes of Pcl mutants, further indicating that RPD3 is required for PcG silencing and possibly for PCL function. These results suggest that there may be multiple ESC/E(Z) complexes with distinct functions in vivo.
Figures
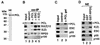

Similar articles
-
The Drosophila Polycomb Group proteins ESC and E(Z) are present in a complex containing the histone-binding protein p55 and the histone deacetylase RPD3.Development. 2001 Jan;128(2):275-86. doi: 10.1242/dev.128.2.275. Development. 2001. PMID: 11124122
-
Polycomblike PHD fingers mediate conserved interaction with enhancer of zeste protein.J Biol Chem. 2001 Nov 16;276(46):43065-73. doi: 10.1074/jbc.M104294200. Epub 2001 Sep 24. J Biol Chem. 2001. PMID: 11571280
-
Histone methyltransferase activity of a Drosophila Polycomb group repressor complex.Cell. 2002 Oct 18;111(2):197-208. doi: 10.1016/s0092-8674(02)00976-5. Cell. 2002. PMID: 12408864
-
The functions of E(Z)/EZH2-mediated methylation of lysine 27 in histone H3.Curr Opin Genet Dev. 2004 Apr;14(2):155-64. doi: 10.1016/j.gde.2004.02.001. Curr Opin Genet Dev. 2004. PMID: 15196462 Review.
-
An EZ mark to miss.Cell Stem Cell. 2008 Dec 4;3(6):577-8. doi: 10.1016/j.stem.2008.11.007. Cell Stem Cell. 2008. PMID: 19041770 Free PMC article. Review.
Cited by
-
Polycomb-like 3 promotes polycomb repressive complex 2 binding to CpG islands and embryonic stem cell self-renewal.PLoS Genet. 2012;8(3):e1002576. doi: 10.1371/journal.pgen.1002576. Epub 2012 Mar 15. PLoS Genet. 2012. PMID: 22438827 Free PMC article.
-
Polycomblike 2 facilitates the recruitment of PRC2 Polycomb group complexes to the inactive X chromosome and to target loci in embryonic stem cells.Development. 2011 Apr;138(8):1471-82. doi: 10.1242/dev.053652. Epub 2011 Mar 2. Development. 2011. PMID: 21367819 Free PMC article.
-
Effects of EZH2 gene on the growth and migration of hepatocellular carcinoma HepG2 cells.Hepatobiliary Surg Nutr. 2013 Apr;2(2):78-83. doi: 10.3978/j.issn.2304-3881.2012.12.12. Hepatobiliary Surg Nutr. 2013. PMID: 24570920 Free PMC article.
-
Genomic distribution of CHD7 on chromatin tracks H3K4 methylation patterns.Genome Res. 2009 Apr;19(4):590-601. doi: 10.1101/gr.086983.108. Epub 2009 Feb 27. Genome Res. 2009. PMID: 19251738 Free PMC article.
-
Functional interaction between the Drosophila knirps short range transcriptional repressor and RPD3 histone deacetylase.J Biol Chem. 2005 Dec 9;280(49):40757-65. doi: 10.1074/jbc.M506819200. Epub 2005 Sep 26. J Biol Chem. 2005. PMID: 16186109 Free PMC article.
References
-
- Aasland, R., T. J. Gibson, and A. F. Stewart. 1995. The PHD finger: implications for chromatin-mediated transcriptional regulation. Trends Biochem. Sci. 20:56-59. - PubMed
-
- Adams, M. D., S. E. Celniker, et al. 2000. The genome sequence of Drosophila melanogaster. Science 287:2185-2195. - PubMed
-
- Anderson, M., K. Fair, S. Amero, S. Nelson, P. J. Harte, and M. O. Diaz. 2002. A new family of cyclophilins with an RNA recognition motif that interact with members of the trx/MLL protein family in Drosophila and human cells. Dev. Genes Evol. 212:107-113. - PubMed
-
- Bajusz, I., L. Sipos, Z. Gyorgypal, E. A. Carrington, R. S. Jones, J. Gausz, and H. Gyurkovics. 2001. The Trithorax-mimic allele of Enhancer of zeste renders active domains of target genes accessible to Polycomb-Group-dependent silencing in Drosophila melanogaster. Genetics 159:1135-1150. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
