Multiple, dispersed human U6 small nuclear RNA genes with varied transcriptional efficiencies
- PMID: 12711679
- PMCID: PMC154217
- DOI: 10.1093/nar/gkg331
Multiple, dispersed human U6 small nuclear RNA genes with varied transcriptional efficiencies
Abstract
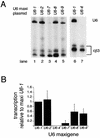
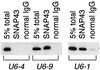
Vertebrate U6 small nuclear RNA (snRNA) gene promoters are among the founding members of those recognized by RNA polymerase III in which all control elements for initiation are located in the 5'-flanking region. Previously, one human U6 gene (U6-1) has been studied extensively. We have identified a total of nine full-length U6 loci in the human genome. Unlike human U1 and U2 snRNA genes, most of the full-length U6 loci are dispersed throughout the genome. Of the nine full-length U6 loci, five are potentially active genes (U6-1, U6-2, U6-7, U6-8 and U6-9) since they are bound by TATA-binding protein and enriched in acetylated histone H4 in cultured human 293 cells. These five all contain OCT, SPH, PSE and TATA elements, although the sequences of these elements are variable. Furthermore, these five genes are transcribed to different extents in vitro or after transient transfection of human 293 cells. Of the nine full-length U6 loci, only U6-7 and U6-8 are closely linked and contain highly conserved 5'-flanking regions. However, due to a modest sequence difference in the proximal sequence elements for U6-7 and U6-8, these genes are transcribed at very different levels in transfected cells.
Figures





Similar articles
-
Zebrafish U6 small nuclear RNA gene promoters contain a SPH element in an unusual location.Gene. 2008 Sep 15;421(1-2):89-94. doi: 10.1016/j.gene.2008.06.019. Epub 2008 Jun 21. Gene. 2008. PMID: 18619527
-
The transcriptional start site for a human U6 small nuclear RNA gene is dictated by a compound promoter element consisting of the PSE and the TATA box.Nucleic Acids Res. 1992 Sep 25;20(18):4903-12. doi: 10.1093/nar/20.18.4903. Nucleic Acids Res. 1992. PMID: 1408805 Free PMC article.
-
U6 snRNA genes of Arabidopsis are transcribed by RNA polymerase III but contain the same two upstream promoter elements as RNA polymerase II-transcribed U-snRNA genes.Nucleic Acids Res. 1990 Jun 25;18(12):3451-8. doi: 10.1093/nar/18.12.3451. Nucleic Acids Res. 1990. PMID: 2362802 Free PMC article.
-
Transcriptional regulation of human small nuclear RNA genes.Biochim Biophys Acta. 2008 May;1779(5):295-305. doi: 10.1016/j.bbagrm.2008.04.001. Epub 2008 Apr 8. Biochim Biophys Acta. 2008. PMID: 18442490 Free PMC article. Review.
-
Structure and expression of U-snRNA genes.Mol Biol Rep. 1987;12(3):139-43. doi: 10.1007/BF00356867. Mol Biol Rep. 1987. PMID: 3316972 Review. No abstract available.
Cited by
-
Structural basis of human SNAPc recognizing proximal sequence element of snRNA promoter.Nat Commun. 2022 Nov 11;13(1):6871. doi: 10.1038/s41467-022-34639-1. Nat Commun. 2022. PMID: 36369505 Free PMC article.
-
U6 snRNA Pseudogenes: Markers of Retrotransposition Dynamics in Mammals.Mol Biol Evol. 2015 Jul;32(7):1815-32. doi: 10.1093/molbev/msv062. Epub 2015 Mar 11. Mol Biol Evol. 2015. PMID: 25761766 Free PMC article.
-
Genomic study of RNA polymerase II and III SNAPc-bound promoters reveals a gene transcribed by both enzymes and a broad use of common activators.PLoS Genet. 2012;8(11):e1003028. doi: 10.1371/journal.pgen.1003028. Epub 2012 Nov 15. PLoS Genet. 2012. PMID: 23166507 Free PMC article.
-
CHD8 associates with human Staf and contributes to efficient U6 RNA polymerase III transcription.Mol Cell Biol. 2007 Dec;27(24):8729-38. doi: 10.1128/MCB.00846-07. Epub 2007 Oct 15. Mol Cell Biol. 2007. PMID: 17938208 Free PMC article.
-
Evolutionary patterns of non-coding RNAs.Theory Biosci. 2005 Apr;123(4):301-69. doi: 10.1016/j.thbio.2005.01.002. Theory Biosci. 2005. PMID: 18202870
References
-
- Hernandez N. (2001) Small nuclear RNA genes: a model system to study fundamental mechanisms of transcription. J. Biol. Chem., 276, 26733–26736. - PubMed
-
- Mattaj I.W., Dathan,N.A., Parry,H.D., Carbon,P. and Krol,A. (1988) Changing the RNA polymerase specificity of U snRNA gene promoters. Cell, 55, 435–442. - PubMed
-
- Lobo S.M. and Hernandez,N. (1989) A 7 bp mutation converts a human RNA polymerase II snRNA promoter into an RNA polymerase III promoter. Cell, 58, 55–67. - PubMed
-
- Kunkel G.R., Cheung,T.C., Miyake,J.H., Urso,O., McNamara-Schroeder,K.J. and Stumph,W.E. (1996) Identification of a SPH element in the distal region of a human U6 small nuclear RNA gene promoter and characterization of the SPH binding factor in HeLa cell extracts. Gene Expr., 6, 59–72. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

